News • Watching the change
Predicting cancer risk with computational electrodynamics
Researchers from Northwestern University are using Argonne supercomputers to advance the development of an optical microscopy technique that can predict and quantify cancer risks at extremely early stages.
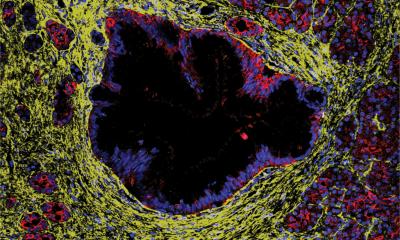
Image courtesy of Vadim Backman, Northwestern University
The basic principle driving Allen Taflove’s computational electrodynamics research — which bears the potential to transform how we diagnose, and possibly treat, various forms of cancer — is strikingly straightforward when distilled: “Simply speaking, we will learn much more about what goes on inside a living cell because we will literally get to watch it change.”
The Northwestern University (NU) professor leads a team of researchers leveraging supercomputing resources at the U.S. Department of Energy’s (DOE) Argonne National Laboratory so as to develop a low-cost, high-throughput optical microscopy technique capable of detecting macromolecular alterations for predicting and quantifying — at extremely early stages — the risk of cancer in humans. More specifically, by running the open-source simulation tool Angora (a “microscope in a computer”) on the Argonne Leadership Computing Facility’s (ALCF) massively parallel Mira system, the team is building upon and moving beyond so-called static partial wave spectroscopy (PWS), which analyzes intracellular activity based on single-instance snapshots, to thereby establish dynamic PWS, which conveys information about a cell’s inner processes by observing that cell’s evolution through time. The ALCF is a DOE Office of Science User Facility.
Recommended article

News • Personalised medicine
Why digital twins could be the ideal therapy testbed
Advanced computer models of diseases can be used to improve diagnosis and treatment. The goal is to develop the models to “digital twins” of individual patients. Those twins may help to computationally identify and try the best medication, before actually treating a patient. The models are the result of an international study, published in the open access journal Genome Medicine.
The "field effect"
In general, PWS analyzes cellular images for certain abnormalities and alterations within these processes — the complex motions and interactions of countless individual molecules — and the intracellular structures to which they are related for signs of early-stage cancer. The relationship between molecular motion and macromolecular structure becomes very difficult to probe at nanometer lengths, which is the relevant scale. By advancing from static analysis to that of live cells, dynamic PWS stands to substantially improve the sensitivity and selectivity of the PWS technique.
Vadim Backman, Walter Dill Scott Professor of Biomedical Engineering at NU and co-principal investigator for the project, was among the first researchers to validate what has been termed the “field effect.” The field effect posits that cancer detection can be accomplished by properly analyzing seemingly normal tissue located some distance from afflicted lesions. If valid, this would create the possibility, for example, that physicians could use buccal swabs to identify lung cancer, or use Pap smears to detect ovarian and cervical cancer. Backman developed the PWS technique precisely to implement such field-effect analyses. By first measuring and then unfolding data for the color spectra of pixels in the backscattering microscope image of a biological cell, he proposed that PWS could determine the degree of randomness of nanometric fluctuations of density within the cell. The degree of randomness, in turn, could be correlated to the presence of a cancer deeper in the body whose “field” includes the cell under analysis.
Backman’s proposal was the source of some controversy given that the intracellular density fluctuations in question — ranging from 20 to 50 nanometers in size — are smaller than the classical diffraction limit at visible wavelengths. Quelling skepticism, in 2013 Backman’s group published in Physical Review Letters (PRL) a rigorous, first-principles theoretical and computational analysis of PWS microscopy (i.e., one derived from Maxwell’s equations). This analysis demonstrated that PWS can, indeed, accurately characterize the statistics of subdiffraction-scale density fluctuations; in fact, the spatial extent of these fluctuations is limited at the low end only by the signal-to-noise ratio of the PWS instrument.
Rigorously solving Maxwell’s equations on nanometer-size voxels ultimately allows Angora to create full-color pixels at the image plane. These pixels can then be analyzed for spectral content
Allen Taflove
The Maxwell’s equations computational solver employed in the 2013 PRL paper was named Angora. Developed over a 10-year period in Backman’s lab under the supervision of Taflove, Angora is built upon a finite-difference time domain (FDTD) physics kernel that solves Maxwell’s equations on nanoscale voxels within the simulated biological cell for an arbitrary lens-focused, apertured illumination. Integral transforms simulating refocusing lenses and apertures are applied to the FDTD-computed optical electromagnetic near field, culminating in the synthesis of true-color microscope images.
“Rigorously solving Maxwell’s equations on nanometer-size voxels ultimately allows Angora to create full-color pixels at the image plane. These pixels can then be analyzed for spectral content,” Taflove said. “After optimizing Angora for Mira, we’ve been able to solve for more than one trillion vector electromagnetic field components, which we think is unique among such software.” Furthermore, by distinguishing between 20-nanometer fluctuations and 50-nanometer fluctuations, Angora far exceeds the capabilities of conventional microscopes.
To date, the Backman group has used PWS to successfully quantify the risk of cancer in seven different organs — lung, colon, pancreas, esophagus, prostate, ovary and thyroid — across several hundred patients. While dynamic PWS has been employed in fewer than ten of these cases so far (the vast majority instead making use of static PWS), the team has plans to assess several thousand more patients so as to generate the reliable statistics needed to gain approval from the U.S. Food and Drug Administration.
Of great interest is a possible next step: building upon the findings of PWS to explore the development of novel cancer therapies. The seven cancers studied with PWS have, historically speaking, been considered disparate diseases, with each demanding a unique, specially tailored treatment. But PWS has shown that unusually heightened intracellular-media density fluctuations are common to all, particularly as regards the chromatin within the nucleus.
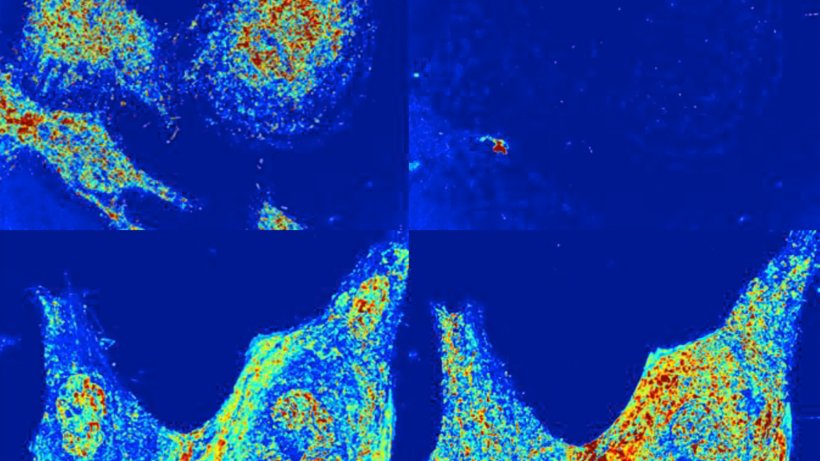
Image courtesy of Vadim Backman, Northwestern University
In light of this discovery, Backman is investigating a potential broad-spectrum cancer treatment he calls chromatin-protection therapy (CPT). Underlying CPT is the hypothesis that if density fluctuations of chromatin within a cancer cell’s nucleus could, via some apposite medication, be reduced to those of a normal cell, then the cancer cell would be more susceptible to deactivation executed through standard treatments such as radiation or chemotherapy. Tests of CPT have already progressed to include a small number of human patients. In support of the CPT hypothesis, Taflove and Backman are proposing a series of Angora FDTD simulations of dynamic PWS for living cancer cells using ALCF supercomputers. “Here, stochastic models for the folding and spatial reorganization of the chromatin within the cell’s nucleus will provide high-spatial-resolution input data to Angora models of dynamic PWS,” Taflove explained.
A paper published in Nature Communications represents the team’s most impactful work yet. Employing dynamic PWS, the researchers measured the intracellular nanoscale structure and macromolecular dynamics of living cells, sensitive to changes as small as 20 nanometers and achieving millisecond temporal resolution. FDTD computations performed with Angora were validated by comparing them with experimental measurements involving nanosphere imaging phantoms. Applying this system in vitro enabled the researchers to explore higher-order chromatin structure and dynamics changes attributable to cellular fixation, stem-cell differentiation, and ultraviolet (UV) irradiation. In doing so, a new phenomenon was discovered: cellular paroxysm, a synchronous, nearly instantaneous burst of intracellular motion that occurs early in the process of UV-induced cell death.
“The combination of our experimental and Angora computational studies has given us confidence that dynamic PWS provides the means with which to obtain nanoscale-sensitive, millisecond-resolved information within living cells without the need for any staining,” said Taflove, who then noted the undesirability of staining techniques, some of which require cytotoxic dyes that can alter cellular behavior. “The impressive spatial-temporal resolution capabilities of dynamic PWS open the door for high-fidelity, high-throughput, early-stage cancer screening, and possibly even novel cancer therapies if the CPT hypothesis is proven.”
Source: Argonne National Laboratory
12.08.2019