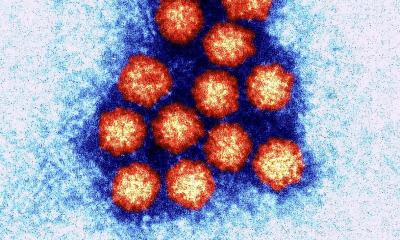
Source: Centers for Disease Control and Prevention
News • Researchers use supercomputer
New insights into hepatitis B
Researchers at the University of Delaware, using supercomputing resources and collaborating with scientists at Indiana University, have gained new understanding of the virus that causes hepatitis B and the "spiky ball" that encloses the virus's genetic blueprint.
The research, which has been published online, ahead of print, by the American Chemical Association journal ACS Chemical Biology, provides insights into how the capsid - a protein shell that protects the blueprint and also drives the delivery of it to infect a host cell -assembles itself.
Computer simulations performed by the UD scientists investigated the effects of a mutation that impairs the assembly process. Together with collaborators, the researchers revealed that the region of the protein that contains the mutation, the spike, can communicate with the region of the protein that links with other subunits to assemble the capsid. They found evidence that a change in the shape of the capsid protein switches it into an "on" state for assembly.
Recommended article

News • Reduced liver complications
The surprising benefits of anti-hepatitis medicine
A new effective treatment of hepatitis C that was made available to all Danes last year, not only combats the virus, but is also effective against potentially fatal complications such as reduced liver functioning and cirrhosis. This is the result of a new study from Aarhus University and Aarhus University Hospital. Hepatitis C is a serious disease, but the biggest threat to someone’s health is…
Scientists believe that the capsid is an important target in developing drugs to treat hepatitis B, a life-threatening and incurable infection that afflicts more than 250 million people worldwide.
"The capsid looks like a spiky ball, with 120 protein dimers that assemble to form it; each dimer contains a spike," said Jodi A. Hadden-Perilla, assistant professor in UD's Department of Chemistry and Biochemistry and a co-author of the new paper. "The capsid is key to the virus infection cycle. If we could disrupt the assembly process, the virus wouldn't be able to produce infectious copies of itself."
The Indiana University researchers had been studying the dimers, which are two-part, T-shaped molecular structures, and investigating whether a mutation could activate or deactivate a switch to turn on the capsid's assembly mechanism. They worked with Hadden-Perilla's group, which ran computer simulations to explain how changes in the protein structure induced by the mutation affected the capsid's ability to assemble.
"What we learned is that this mutation disrupts the structure of the spike at the top of the dimer," Hadden-Perilla said. "This mutation slows down assembly, which actually involves a region of the protein that is far away from the spike. It's clear that these two regions are connected. A change in the shape of the protein, particularly at the spike, may actually activate or deactivate assembly."
Her team did its work using the National Science Foundation-supported Blue Waters supercomputer at the University of Illinois at Urbana-Champaign, the largest supercomputer on any university campus in the world, to perform what are known as all-atom molecular dynamics simulations.
Molecular dynamics simulations allow researchers to study the way molecules move in order to learn how they carry out their functions in nature. Computer simulations are the only method that can reveal the motion of molecular systems down to the atomic level and are sometimes referred to as the "computational microscope."
Source: University of Delaware
16.08.2020