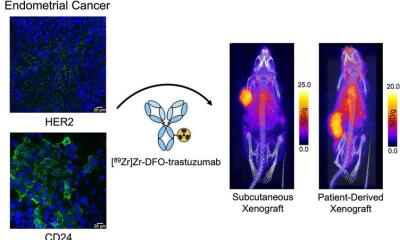
Interview • Sequencing
Precision medicine in oncology
Professor Hedvig Hricak MD PhD, Chair of the Department of Radiology at the Memorial Sloan-Kettering Cancer Centre, New York, and Professor of Radiology at Cornell University, Ithaca, New York, is a notable expert on crosssectional anatomic and molecular imaging, particularly of gynaecologic and prostate cancers. EH interviewed her about the potential and impact of more precise viewing of inter- and intra-tumoral heterogeneity.
Interview: John Brosky
Precision medicine in oncology offers tremendous potential and challenges. Are there obstacles?

‘Probably one of the greatest challenges to the implementation of precision medicine in oncology is the tremendous inter- and intra-tumoral genetic heterogeneity. The notions of inter- and intra-tumour heterogeneity have been recognised for many years, but recent advances in sequencing technology are allowing the true extent of both forms of heterogeneity to be revealed in detail.
Furthermore, sequential analysis of tumours has also revealed that intra-tumour heterogeneity temporally evolves during the disease course, which enables development of tumour resistance to therapies. Imaging has a tremendous potential to play in addressing this challenge, as it is the only tool that can examine biological heterogeneity both within an entire tumour and across multiple tumour deposits in the body, in vivo.
However, wide-spread use of imaging for this purpose will require tremendous further research, test-validation further combine with overcoming regulatory hurdles to tracer development and approval, educating current and future radiologists to have a deeper understanding of molecular biology, and implementing bioinformatics on a much larger scale.’
You have said MRI has extensive untapped potential to contribute to molecularly-based precision medicine. Where is this being realised?
‘I’d like to point out that my comments are limited to the abdominal and pelvic MR – as those are the areas of my expertise. Functional MRI techniques, such as dynamic contrast-enhanced (DCE) and diffusion-weighted (DW) MRI, are already providing prognostic, predictive, and early-response biomarkers that can be used to help determine the need for treatment, predict treatment response and assess the effectiveness of treatment early on.
However, many of these biomarkers still need to be validated in multicentre prospective studies. In the future, the new discipline of ‘radiomics’ promises to dramatically expand the number of imaging biomarkers we can derive from MRI. With radiomics analyses, a large number of features quantifying tumour signal intensity, texture and shape, as well as functional parameters and clusters of multiparametric data, can be extracted from MR images and correlated with treatment outcome data.
Furthermore, in an approach called ‘radiogenomics’, such features can be correlated with omic data, including specific gene clusters. By providing spatially and temporally specific information about tumour biology, radiomics and radiogenomics will enable radiologists to recommend where to biopsy, make predictions about tumour aggressiveness in areas where biopsy is not feasible, and improve treatment selection and planning as well as assessment of treatment response. As discussed (see below), hyperpolarised MRSI is another MRI technique with great potential to contribute to precision medicine.’
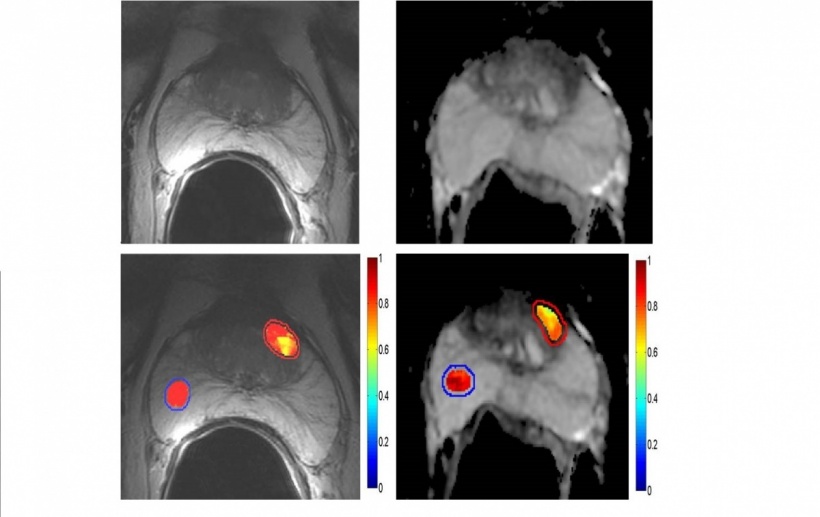
Tumor Detection/Aggressiveness.
Texture Analysis Compared to T2WI & DWI there is further improvement in PCa detection, visualization of tumor heterogeneity and
tumor characterization
Hyperpolarised MR spectroscopic imaging may revolutionise the way MRI is used in cancer care. What does this revolution look like?
‘Hyperpolarised MR spectroscopic imaging (HP-MRSI) is a new technology that increases the MR signal 10,000–100,000-fold, and therefore enables MR imaging of nuclei other than 1H with great speed and sensitivity. Imaging after injection of a hyperpolarised agent, such as 13C-pyruvate, allows visualisation of the distribution of the agent itself as well as its downstream enzymatic products. By allowing precise identification of aberrant molecular processes, HP-MRSI should enable better treatment selection and earlier assessment of treatment response.’
How do you see these advanced MRI techniques being translated into clinical routine for greater precision in medicine?
‘Importantly, HP-MRSI allows short imaging times (seconds to minutes) that can be added to existing protocols without significantly affecting workflow, and injected HP-MRSI agents are naturally occurring substances with no inherent toxicity, making them safe for use in patients. Because of these practical characteristics, HP-MRSI could easily be incorporated into routine MRI examinations that include other sequences, such as T-2-weighted imaging, DCE-MRI or DW-MRI.
When these capabilities are also combined with an overlay of augmented information from radiomics and radiogenomics, MRI may become an extremely powerful tool for increasing precision in all areas of cancer care, from diagnosis to treatment selection and planning, treatment monitoring and follow up. Of note, machine learning, construction of radiomics algorithms and automated pattern recognition should make it possible to develop augmented programmes and therefore disseminate and introduce the added value of radiomics/radiogenomics in clinical practice, thus improving accuracy in oncologic imaging among radiologists who do not sub-specialise in the field. Getting to that point will need a great deal of teamwork and much greater integration of advanced biomedical informatics in clinical settings.’
07.04.2015