Image source: Bruker
Article • Infection control
The strain typing technologies of tomorrow
Cedars-Sinai Medical Center, a non-profit hospital and medical research institution in Los Angeles, is setting new standards for quality and innovation in patient care by successfully introducing typing of Candida auris species – a procedure that could prove crucial in protecting patients from infection outbreaks caused by these microbes in healthcare settings.
Authors:
Dr. Margie Morgan, Medical Director of Clinical Microbiology for Pathology and Laboratory Medicine at Cedars-Sinai Medical Center
Dr. Deisy Contreras, Clinical Associate of the Clinical Microbiology Laboratory in the Department of Pathology and Laboratory Medicine at Cedars-Sinai
Dr. Markus Meyer, Head of Business Unit Hygiene & Epidemiology, Bruker Microbiology & Infection Diagnostics
The first reports of C. auris in the US came in 2015, but ongoing local transmission has since led to over 1,000 cumulative cases in Los Angeles County.1,2 Infection with C. auris is associated with high mortality rates, and it is often resistant to multiple classes of antifungal drugs.3 It spreads easily in hospital environments via colonized patients and contaminated surfaces or equipment.
The first step in fighting these infections is to identify the microbe responsible. Fast species identification followed by strain typing – to reveal clonal relations or differentiate genetic lineages – is beneficial. Traditional strain typing technologies like pulsed-field gel electrophoresis, multi-locus sequence typing, latex agglutination, and whole-genome sequencing, however, are time-consuming and resource-intensive. These technologies are also not readily available in all microbiology labs, which raises the question: could alternative emerging typing technologies fill the void?
New strain typing technologies
Microorganism strain typing is vital for infection control. Modern strain typing strategies tend towards molecular fingerprinting technologies.4 Fourier transform infrared (FT-IR) spectroscopy has shown promise as a rapid and non-invasive tool that requires minimal training to master. This technology delivers rapid strain-level discrimination of microbes, starting from culture, to provide a simpler alternative to next-generation sequencing strain typing.
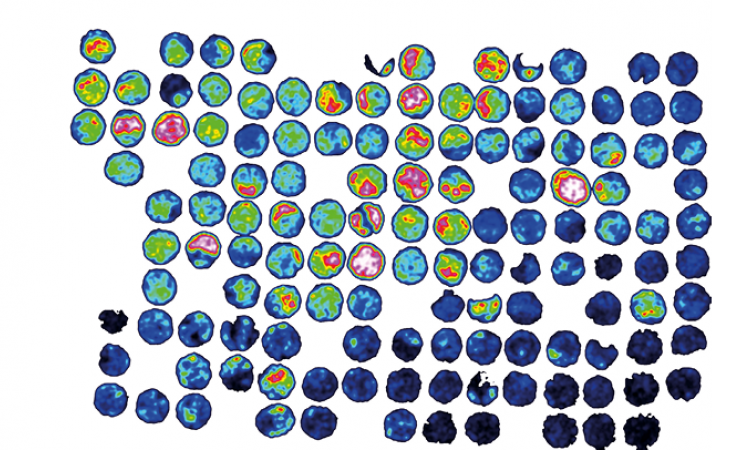
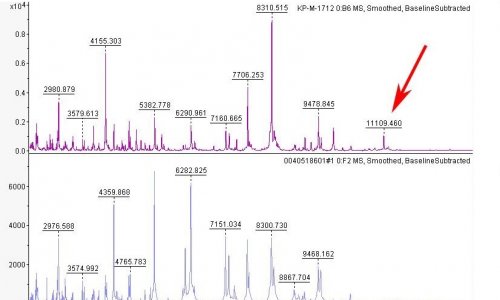
IR spectroscopy measures the molecular vibrations associated with the absorption of IR light. Different chemical structures vibrate at different frequencies when this absorption happens. For example, the carboxyl group in fatty acids and lipids vibrates at 2800-3000 cm-1 and the amide group in proteins vibrates at 1500-1800 cm-1. IR-based strain typing works by combining the information from these absorption ranges to produce a molecular fingerprint for a given sample. Microorganisms can be identified through the recognition of motifs in the fingerprint, particularly those belonging to carbohydrates. IR-based strain typing is relatively simple when compared with more traditional methods and achieves results faster, taking just 30 minutes for a single sample.
Researchers at Cedars-Sinai are capitalizing on the benefits of these technologies to find strategies to combat the threat presented by C. auris.
Targeting C. auris
Work conducted at Cedars-Sinai has helped guide the way in clinical applications for IR spectroscopy in California. The institute was the first in the US to evaluate a newly available FT-IR spectroscopy system for strain typing. Initial work focused on outbreaks of Gram-negative rod bacteria (such as Pseudomonas aeruginosa) and Gram-positive cocci (like Staphylococcus aureus), but the team soon turned its attention to C. auris. By 2021, IR spectroscopy was well-established for routine C. auris testing at Cedars-Sinai.
When the pandemic hit, the team sought to find a way to conduct surveillance for C. auris and prevent its transmission to at-risk individuals before admission to hospital. The initial real-time polymerase chain reaction (RT-PCR) platform for C. auris identification worked, but the hospital wanted to implement another layer of surveillance – and turned to IR technology.
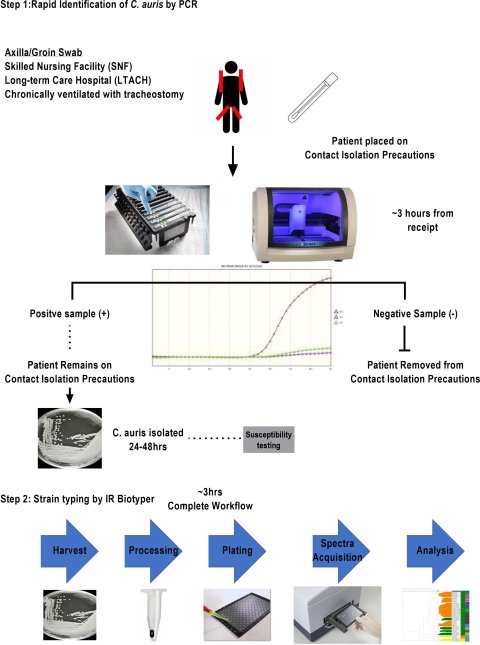
Image source: Contreras DA, Morgan MA, Frontiers in Cellular and Infection Microbiology 2022 (CC BY 4.0)
Today, researchers at Cedars-Sinai are pioneering a first line of defense system. The first step is to detect C. auris via axilla or groin swabs using PCR to identify at-risk patients quickly and place them under the appropriate contact prevention precautions. The second step is to identify the strain type of the fungus via IR spectroscopy, starting from culture, for positive samples.
Identifying C.auris with PCR can provide an early warning system and help guide clinical decisions at the early stages of infection. Speed at this stage is critical to improve patient outcomes and, so far, the results have been promising. The researchers found that 4% of more than 700 at-risk patients (28 patients) at the center tested positive for C. auris,5 with low numbers of genomic variation indicating local and ongoing transmission within the Los Angeles area, not exclusively within the hospital setting.
The institute has built a database of everyone admitted to Cedars-Sinai who tested positive for C. auris. This valuable resource will help to identify possible future outbreaks rapidly.
Looking forward
Antimicrobial resistance (AMR) is considered one of the greatest health threats facing humanity.6 Despite the rapid global spread, it is difficult to predict the actual burden of the infection as the standard laboratory methods fail to correctly identify the fungi.3 This emphasizes the importance of the strain typing work that is ongoing at Cedars-Sinai, and targeting C. auris is only the beginning. The Cedars-Sinai team is now evaluating IR spectroscopy for the identification of Mycobacterium abscessus subspecies. Validation of this application could facilitate strain typing and clarithromycin susceptibility testing for Mycobacterium abscessus in a shorter time than current methods.
What factors will ensure the success of tomorrow’s typing technologies? Thinking outside of the box, innovative instrumentation, and clear communication are among the most important qualities to consider. Central to this will be increasing awareness and advancing the diagnostic methods for early disease detection and control.
Note: The testing procedure was developed by the researchers for the specific use in the Cedars-Sinai Medical Center only. The IR Biotyper is not intended for the examination of specimen from human body to define or monitor therapeutic measures.
Profiles:
Dr. Margie Morgan is the Medical Director of Clinical Microbiology for Pathology and Laboratory Medicine at Cedars-Sinai Medical Center. Dr. Morgan oversees clinical testing within the microbiology laboratory, guides staff conducting diagnostic tests, provides teaching services for residents and fellows, and participates in researching new diagnostic techniques within microbiology.
Dr. Deisy Contreras is the Clinical Associate of the Clinical Microbiology Laboratory in the Department of Pathology and Laboratory Medicine at Cedars-Sinai. Dr. Contreras researches the application of diagnostic techniques within laboratory workflows.
Markus Meyer, PhD, is head of the Business Unit Hygiene/Epidemiology at the Microbiology and Infection Diagnostics (MID) Division of Bruker Daltonics GmbH & Co. KG in Bremen, Germany. He has been with Bruker Daltonics since 2010 as a product manager for various life science mass spectrometry instruments before joining the Microbiology & Infection Diagnostics team as global product manager for consumables in 2018.
References:
- Chow NA et al.: Multiple introductions and subsequent transmission of multidrug-resistant Candida auris in the USA: a molecular epidemiological survey; Lancet Infectious Diseases 2018
- County of Los Angeles Public Health: Mitigating the Spread of Candida auris in Los Angeles Country
- Ademe M et al.: Candida auris: From Multidrug Resistance to Pan-Resistant Strains; Infection and Drug Resistance 2020
- Franco-Duarte R et al.: Advances in Chemical and Biological Methods to Identify Microorganisms-From Past to Present; Microorganisms 2019
- Contreras DA et al.: Surveillance diagnostic algorithm using real-time PCR assay and strain typing method development to assist with the control of C. auris amid COVID-19 pandemic; Frontiers in Cellular and Infection Microbiology 2022
- World Health Organization: Antimicrobial resistance 2021 (Accessed April 2023)
20.11.2023
- bacteria (243)
- diagnostics (584)
- hospital (321)
- infections (770)
- laboratory (1124)
- microbiology (295)
- pathology (299)
- prevention (707)