News • Gene editing
Taking CRISPR one step further
Researchers at ETH Zurich have refined the famous CRISPR-Cas method. Now, for the very first time, it is possible to modify dozens, if not hundreds, of genes in a cell simultaneously.
The biotechnological method CRISPR-Cas offers a relatively quick and easy way to manipulate single genes in cells, meaning they can be precisely deleted, replaced or modified. Furthermore, in recent years, researchers have also been using technologies based on CRISPR-Cas to systematically increase or decrease the activity of individual genes. The corresponding methods have become the worldwide standard within a very short time, both in basic biological research and in applied fields such as plant breeding.
Recommended article
News • Gene editing
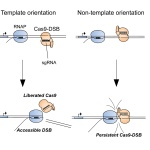
Why CRISPR fails sometimes – and what to do about it
CRISPR has recently become synonymous with precise genetic editing. However, the mighty tool has been known to fail from time to time. Now, researchers have found out why.
Our method enables us, for the first time, to systematically modify entire gene networks in a single step
Randall Platt
To date, for the most part, researchers could modify only one gene at a time using the method. On occasion, they managed two or three in one go; in one particular case, they were able to edit seven genes simultaneously. Now, Professor Randall Platt and his team at the Department of Biosystems Science and Engineering at ETH Zurich in Basel have developed a process that – as they demonstrated in experiments – can modify 25 target sites within genes in a cell at once. As if that were not enough, this number can be increased still further, to dozens or even hundreds of genes, as Platt points out. At any rate, the method offers enormous potential for biomedical research and biotechnology. “Thanks to this new tool, we and other scientists can now achieve what we could only dream of doing in the past.”
The researchers present their findings in the journal Nature Methods.
Genes and proteins in cells interact in many different ways. The resulting networks comprising dozens of genes ensure an organism’s cellular diversity. For example, they are responsible for differentiating progenitor cells to neuronal cells and immune cells. “Our method enables us, for the first time, to systematically modify entire gene networks in a single step,” Platt says.
Moreover, it paves the way for complex, large-scale cell programming. It can be used to increase the activity of certain genes, while reducing that of others. The timing of this change in activity can also be precisely controlled. This is of interest for basic research, for example in investigating why various types of cells behave differently or for the study of complex genetic disorders. It will also prove useful for cell replacement therapy, which involves replacing damaged with healthy cells. In this case, researchers can use the method to convert stem cells into differentiated cells, such as neuronal cells or insulin-producing beta cells, or vice versa, to produce stem cells from differentiated skin cells.

Visualizations: ETH Zurich / Carlo Cosimo Campa
The dual function of the Cas enzyme
The CRISPR-Cas method requires an enzyme known as a Cas and a small RNA molecule. Its sequence of nucleobases serves as an “address label”, directing the enzyme with utmost precision to its designated site of action on the chromosomes. ETH scientists have created a plasmid, or a circular DNA molecule, that stores the blueprint of the Cas enzyme and numerous RNA address molecules, arranged in sequences: in other words, a longer address list. In their experiments, the researchers inserted this plasmid into human cells, thereby demonstrating that several genes can be modified and regulated simultaneously.
For the new technique, the scientists did not use the Cas9 enzyme that has featured in most CRISPR-Cas methods to date, but the related Cas12a enzyme. Not only can it edit genes, it can also cut the long “RNA address list” into individual “address labels” at the same time. Furthermore, Cas12a can handle shorter RNA address molecules than Cas9. “The shorter these addressing sequences are, the more of them we can fit onto a plasmid,” Platt says.
Source: ETH Zurich
14.08.2019





