Article • Infections
Carbapenem resistant strains
The increasing numbers of bacteria resistant to the newer generations of antibiotics is a public health problem on a global scale. Bacteria have an extraordinary capacity for adaptation, mutating permanently to overcome the action of our increasingly impotent antimicrobial armamentarium. A situation further aggravated by the use of the powerful ‘large spectrum’ antibiotics, creating further pressure towards acquired resistance.
Report: Jane MacDougall
Source: Shutterstock.com/Aunt Spray
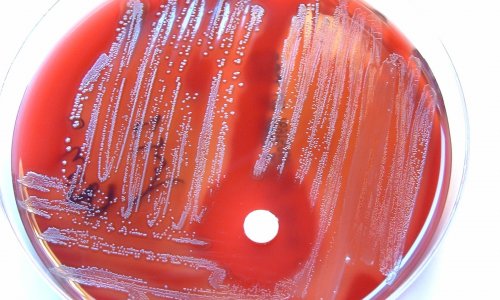
Extended-spectrum beta-lactamases (ESBL) are enzymes that confer resistance to most beta-lactam antibiotics; these include penicillins, cephalosporins and aztreonam. Community and hospital-acquired ESBL-producing Enterobacteriaceae are prevalent worldwide and include common gut microflora, such as Escherichia coli. Infections with ESBL-producing organisms are associated with poor outcome and high mortality rates. However, because the reliable identification of ESBL-producing organisms is challenging, their prevalence is likely to be underestimated.
The first ESBL-producing organisms were isolated in the 1980s; today there are hundreds of different ESBL-producing strains resulting from successive mutations. The emergence, particularly in hospital, of resistant E. coli creates a vicious circle; the use of large spectrum antibiotics essentially leads to the emergence of greater numbers of resistant bacteria.
Christian Cattoen, from the Valenciennes Hospital Group in France, explained how this occurs ‘… the antibiotics are excreted from the body via the liver, therefore arriving in the digestive tract in bile. The normal gut microflora sensitive to the antibiotics is wiped out and in the void that is left, the resistant strains of E. coli, even if only present in very low numbers can now rapidly multiply. Nature abhors a vacuum!’
To combat these bacteria a family of antibiotics, now nearly 20 years old was developed – the carbapenems. The original molecule with anti-b-lactamase activity was derived from Gram-positive bacterium Streptomyces clavuligerus. However, by the 2000s the first carbapenem resistant strains had been isolated. Resistance to carbapenems is now appearing at regular intervals worldwide from different mutations. The discovery of new producers of carbapenemases, a form of b-lactamase, is a worrying trend. ‘While still highly effective against ESBL-producing organisms and other multi drug resistant organisms, both Gram-negative and Gram-positive, the use of these antibiotics must only be considered as a last resort,’ Cattoen insisted.
In 2015, the World Health Organisation issued a warning about the spread of bacteria resistant to carbapenems, drawing attention to the fact that they were now a global phenomenon.
This rapid increase and spread of dangerous bacteria turns the spotlight on the clinical laboratory. Firstly, in helping to reduce the use of broad-spectrum antibiotics by concentrating on new methods for returning an antibiogram in a timely manner, so the clinician can initiate appropriate treatment for the isolated pathogen. Recently developed rapid antimicrobial susceptibility testing methods include classical agglutination assays, molecular testing methods, e.g. qPCR, DNA microarrays, Luminex xMAP assays and next generation sequencing. Additionally, there are fluorescence in situ hybridisation (FISH) and mass spectrometry-based methods, e.g. phyloproteomics, assays using stable isotope labelling of amino acids, mass spectrometric beta-lactamase assays, PCR/electrospray ionisation-mass spectrometry (PCR/ESI MS), mini-sequencing and mass spectrometry-based comparative sequence analysis (MSCSA) all aiming to rapidly identify and determine the susceptibility profile of pathogens.
While, not immediately available to all laboratories, clinical laboratories should certainly think about incorporating techniques, such as DNA chips, to more easily identify carbapenem resistant bacteria, so that infected patients can be quickly isolated and the hospital’s contingency plan for containment implemented. ‘The clinical biologist should be fully implicated in this procedure,’ Cattoen advised. The reliability of our results is primordial, the better the techniques we have, the sooner we can raise the alert and put processes in action, that the clinical laboratory needs to be at the heart of.’
27.10.2016