Article • Mass spectrometry-based metabolomics
A new technique to understand metabolic pathways
Mass spectrometry-based metabolomics has emerged as a powerful tool to help study chemical ecology. Recent advances in the technique make it possible to study microbial interactions from complex communities.
Report: Mark Nicholls
Laia Castaño-Espriu outlined the role and benefits of MS in this context in her presentation ‘Analysis of microbial ecology by mass spectrometry-based metabolomics techniques’, at the joint meeting of the German Society for Hygiene and Microbiology (DGHM) and the Association for General and Applied Microbiology (VAAM) in Leipzig, Germany this March.
Microbial competition enhances chemical diversity

Castaño-Espriu explained that the bacterial order Actinobacteria is responsible for the production of 65-70% of microbially-produced specialised metabolites with diverse biological activities, but it is estimated that only 10% of biosynthetic genes clusters encoding the production of these specialised metabolites are transcribed under normal laboratory conditions. ‘It’s been observed that microbial competition enhances chemical diversity as an ecological advantage,’ she said. ‘Therefore, the study of bacterial competition is key in understanding the ecological role that secondary metabolites pose to microbes and in understanding the induction of cryptic biosynthetic gene clusters.’
As a final year PhD student at the University of Strathclyde in Glasgow, Castaño-Espriu’s project is based on the analysis of microbial interactions by co-cultivation and MS-based metabolomics, with a particular focus on Actinobacteria that has been isolated from the marine environment as a potential source of novel chemistry.
The analysis of these interactions by LC-MS revealed the production of metabolites that were co-culture specific and they found that several interactions not only enhanced chemical diversity, but also inhibited the growth of several clinically relevant pathogens, she explained. ‘Furthermore, we could gain a better understanding of the chemical ecology during bacterial competition, where changing conditions aggravate the complexity of these interactions. Therefore, we demonstrated that MS-based metabolomics is an exciting strategy to study microbial ecology and to prioritise novel chemistry.’
Recommended article
Article • Human-bacterial symbiosis
The value of the human microbiome
The human species maintains symbiotic relationships with a multitude of microbial organisms that colonise the inside as well as the surface of the body. Scientists, for a long time, underestimated the significance of these organisms for humans.
Metabolomics approaches – referring to the study of the complete set of metabolites found in a cell, tissue, organ or organism - in general are applied to compare the chemical profiles of different experimental groups to find out the chemical response to external stimuli such as environmental stresses, she explained. ‘MS-based metabolomics approaches in microbial ecology enable the understanding of microbial interactions in complex communities. Metabolomics allows the analysis of those metabolites that are produced during microbial interactions, therefore, providing a snapshot of the ecological role of these metabolites in the producers. This allows the understanding of biological questions such as "Why are certain metabolites produced under specific conditions?"’, she pointed out.
By studying the secondary metabolism of organisms in complex communities and understanding the role of interesting metabolites, Castaño-Espriu continued, it is possible to gain greater insights in the correlation between phenotypic traits and microbial interactions.
Identification of potentially significant metabolites
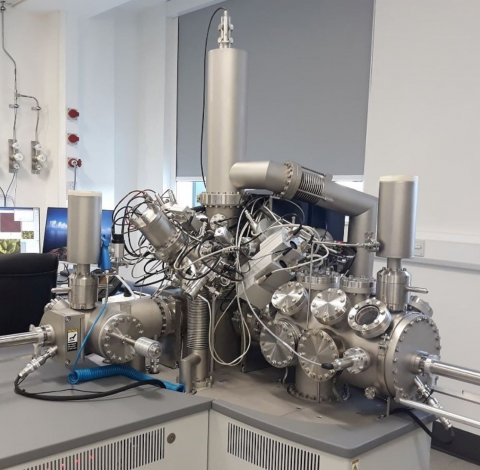

Her work utilises Time-of-Flight Secondary Ion Mass Spectrometry (ToF-SIMS) and Liquid Chromatography Mass Spectrometry (LC-MS). The MS-based metabolomics studies follow the steps of sample collection and preparation, generation and acquisition of MS data (metabolic profiles), data processing using a deconvolution software such as MZmine, and statistical analyses. ‘This approach,’ she said, ‘enables the identification of potentially significant metabolites belonging to the groups of interest, therefore, prioritising these for chemical identification and further experiments to understand the underlying mechanisms of the metabolic pathways.’
MS-based metabolomics techniques offer advantages in fields including the effect of drugs at a metabolite level; drug development; the study of diseases such as cancer, nutrition science, toxicology analyses to natural products; and chemical ecology research. ‘Therefore,’ Castaño-Espriu added, ‘MS-metabolomics represents a multi- and interdisciplinary approach that can be used in different fields to understand metabolic pathways.’
Metabolomics, she added, aims to deliver more rapid and reliable analyses combined with more economical methods, data standardisation and the sharing of more MS data through online databases. Recent applications of metabolomics include disease biomarkers and drug development, with a significant increase in the use of MS-based metabolomics observed in the area of microbial ecology in the last decade.
Compared to other techniques, Castaño-Espriu believes that metabolomics is the best approach to accelerate drug development as it is low-costs and also rapid and effective. ‘This effectiveness makes a metabolomics approach ideal for other life and medical sciences.’ She feels it will become necessary to develop new technologies that offer greater sensitivity and spatial resolution to improve data quality, and to create databases with more MS data coverage to facilitate metabolite identification. Another challenge lies in data variability based on sample preparation and the sample matrix among other factors. However, she believes the development of new instruments and new methods will aim to overcome this limitation.
Profile:
Laia Castaño-Espriu is a final year PhD student at the University of Strathclyde in Glasgow under the supervision of Dr Katherine R. Duncan, with a particular research interest in metabolomics, actinobacteria and drug discovery.
21.10.2020





