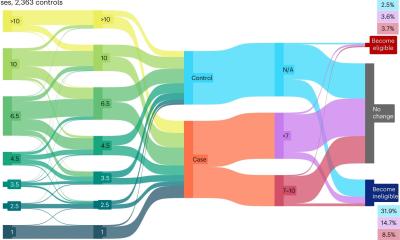
Image source: Montagud et al., eLife 2022 (CC BY 4.0); montage: HiE/Behrends
News • Calculating cancer
Virtual patient ‘surrogates’ to personalise cancer treatments
Scientists have developed mathematical models that act as patient ‘surrogates’ for evaluating potential prostate cancer treatments.
The research, published in eLife, could ultimately help clinicians choose the most effective drug combination before they start to treat a patient, potentially improving their response and avoiding drug resistance.
Researchers used an approach called Boolean modelling, which is already used to describe dynamics of complex cell signalling processes. But existing models have been generic and have not accounted for the differences between individual patients’ diseases or how they respond to treatment. “The dream has always been to use more and more complex models and data until we can have digital twins, or virtual humans or surrogates – a simulation that helps select the proper clinical treatment for a given patient with high degrees of specificity or sensitivity,” explains Arnau Montagud, who was a researcher at Institut Curie, Paris, France, at the time the study was carried out, and is now at the Barcelona Supercomputing Center (BSC), Spain. “We wanted to know if our method of tailoring Boolean models of cell signalling was accurate enough to discriminate between different patients, and whether the models could be used as testbeds to rank personalised drug treatments.”
To begin, the team used data from The Cancer Genome Atlas (TCGA) and other databases to create a network of all relevant pathways involved in prostate cell signalling. Then they converted this into a generic Boolean model where all the nodes in the network can be assigned one of two values – 0 (inactivated or absent) or 1 (activated or present). Data from 488 prostate cancer patients from TCGA were used to create 488 patient-specific Boolean models. For example, where a patient’s tumour had a mutation in a specific gene, this meant the node in the network was inactivated, and assigned a value of 0.
Recommended article
Article • Datasets
Benefits from The Cancer Genome Atlas
Last year, scientists at the University of California San Francisco (UCSF) revealed that by measuring the proportion of both immune and cancerous cells in tumours, or ‘tumour purity,’ clinicians could more precisely predict the success of certain precision therapies. A key aspect of the discovery was access to over 10,000 samples constituting 21 different cancers.
Having built these models, the team looked in each patient model for genes that, when inhibited, would block growth or encourage death of cancer cells. They narrowed these genes down to a list of targets of existing drugs, and ran simulations to predict what would happen if the drugs were combined. This allowed them to compare the effects of individual drugs on each patient, and to propose certain drugs that would work for specific patients or for groups of patients. Inactivation of some of the genes had a greater effect in some patients compared with others, highlighting opportunities for personalised drug treatments. The simulations also spotted patterns linked to the grade of patients’ tumours as measured by the Gleason score, suggesting it might be possible to tailor drug treatments to prostate cancer patients according to their score in the future.
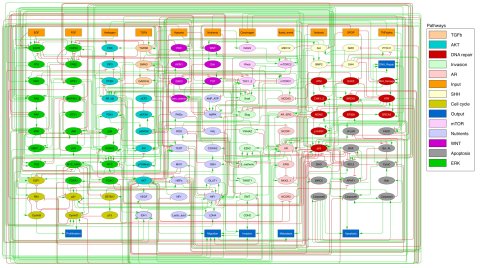
Image source: Montagud et al., eLife 2022 (CC BY 4.0)
Testing whether these treatment predictions hold true in patients would require a clinical trial, so the team instead built eight different personalised prostate cancer cell line models from publicly available data. As with the patient models, they looked for commonly occurring mutations in the cell lines that influenced cancer cell growth or death. This resulted in the identification of 17 proteins that could be targeted with drugs.
Next, to investigate if drugs targeting these proteins would have the anticipated effects, they mimicked the effect of different drug dosages in the model by switching off each node from 100% active to 0% active and looking at the effects on growth, death and spread of the cancer cells. When they carried out the same experiment in real cell lines, it confirmed that blocking the identified nodes in the model had differential effects on cell growth and survival. Moreover, the model could predict synergistic effects of treatments that work against different nodes in the network, which could help to identify promising drug combinations for future investigation. “Our personalised models suggest single and combined drug treatments for individual prostate cancer patients,” concludes Laurence Calzone, a researcher at Institut Curie, and a co-senior author of the study alongside Julio Saez-Rodriguez, from Heidelberg University, Germany. “These advances are incremental steps towards having digital twins that will help clinicians before they go to the patient's bedside, allowing them to capture patient individualities and test and rank different drug treatments.”
Source: eLife
16.02.2022