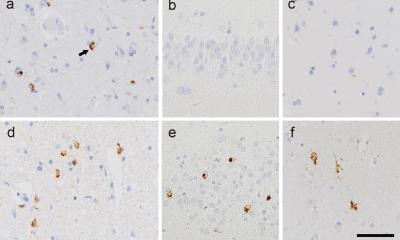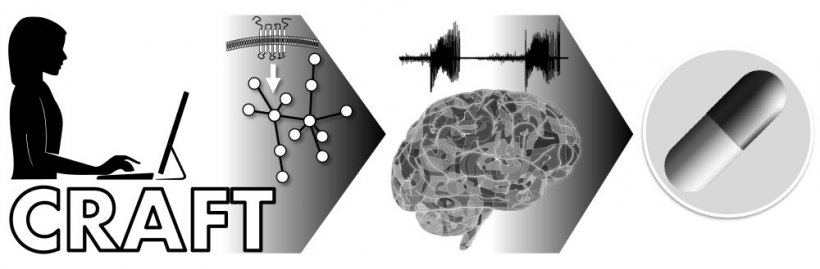
Credit: Enrico Petretto
News • Debilitating disease
Discovery of a new anti-epileptic drug target
An international collaboration between Imperial College London, Singapore's Duke-NUS Medical School and Belgium-based pharmaceutical company UCB led to the discovery of a new anti-epileptic drug target and a whole new approach that promises to speed up the discovery of future drugs to treat debilitating diseases, such as epilepsy.
"The identification of drug targets is highly challenging."
Prof. Michael Johnson
Led by Prof. Michael Johnson, Division of Brain Sciences, Imperial College London, UK; A/Prof. Enrico Petretto, Centre for Computational Biology, Duke-NUS Medical School; and Dr. Rafal Kaminski, UCB, Belgium, this collaboration developed an advanced computational approach to predict new drug targets. As proof-of-concept, the investigators applied their computational approach to epilepsy resulting in the discovery of a new drug target for the disease that was experimentally confirmed by showing that pharmacological blockade of the target had anti-seizure effects in pre-clinical models.
"The identification of drug targets is highly challenging, particularly for diseases of the brain," Prof. Johnson explained. "With our approach – which we named the 'Causal Reasoning Analytical Framework for Target discovery,' or CRAFT – we discovered and validated a potential new anti-epileptic drug in less than two years. This international collaborative study is an example of a successful academia-industry partnership, resulting not only in the discovery of a new drug candidate, but also in an innovation with the promise to transform medicine and improve lives by speeding up the identification of new drug targets across the spectrum of human disease."
Big data
CRAFT draws on genomic 'big data' and applies a systems-level computational framework to drug target discovery that combines gene regulatory information with causal reasoning. Starting from gene expression data from the target tissue, CRAFT's predictive framework identifies cell membrane receptors that play a regulatory role in disease-related gene expression. This enables researchers to understand the mechanism of action of a disease, and computationally predict the effectiveness of a potential drug target.
Epilepsy is a debilitating brain disease for which there is a global unmet need – approximately one in three epilepsy patients are resistant to all currently available anti-epileptic drugs and none of the current drugs are disease modifying or curative. Conventional drug development methods, particularly for diseases of the central nervous system, suffers from a high rate of attrition due to inadequate drug target validation in the early stages of discovery.
"In contrast with traditional drug discovery pipelines, CRAFT provides an efficient data-driven approach based on a systems genetics framework that permits the identification of gene networks driving disease and their master control points in record time – a strategy that we implemented here for predicting membrane receptors as effective drug targets that can then be experimentally validated at the earliest stage of the drug discovery process," explained A/Prof. Petretto who leads the Systems Genetics group at Duke-NUS.
Partnership between academia and industry
"Our collaboration has paved a completely new way of drug discovery in epilepsy."
Dr. Rafal Kaminski
CRAFT's systems genetics approach replaced the traditional approach of examining only one component of a complex system at a time. Prof. Johnson explained: "We first describe the disease in terms of its gene expression signature and then using knowledge of how genes are controlled, CRAFT identifies membrane receptors predicted to exert a regulatory effect over the disease state. We specifically chose to develop a method connecting disease states to druggable membrane receptors because over half of existing drugs already target membrane receptors, and so CRAFT allows the maximum opportunity for drug repurposing and rapid experimental medicine proofs of concept, as well as new drug development. In the case of epilepsy, this led to the identification of the microglial membrane receptor Csf1R as a potential new therapeutic target for epilepsy."
The partnership between academia and industry was pivotal for this discovery. "Our collaboration has paved a completely new way of drug discovery in epilepsy," said Dr. Kaminski from UCB. "We moved away from traditional drug screening approaches into computational identification of key disease drivers in order to match them with already existing drugs that have the desired mode of action. This new strategy has the potential to dramatically accelerate the drug development process and bring new treatments to patients much faster."
Source: Duke-NUS Medical School
03.09.2018