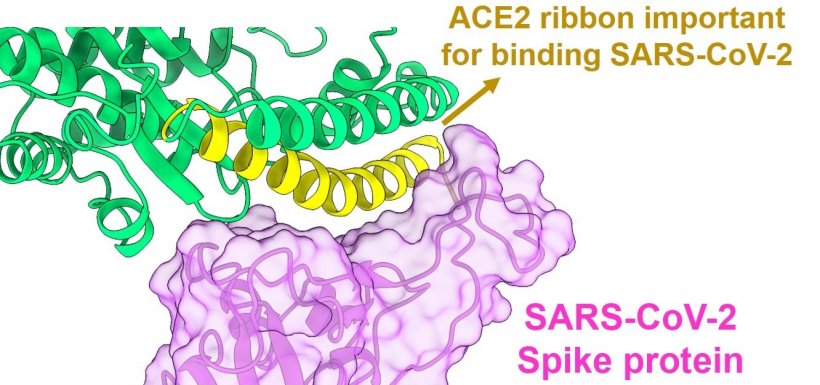
Image source: Ohio State University; graphic generated using ChimeraX software and published protein structure
News • Peptide research
A 'fake handshake' to trick the coronavirus
Scientists have developed peptides that fit snugly into a groove on the SARS-CoV-2 Spike protein that it would normally use to access a host cell. These protein fragments effectively trick the virus into “shaking hands” with a replica rather than with the actual protein on a cell’s surface that lets the virus in.
Previous research has determined that the novel coronavirus binds to a receptor protein on a target cell’s surface called ACE2. This receptor is located on certain types of human cells in the lung and nasal cavity, providing SARS-CoV-2 many access points to infect the body. For this work, Ohio State University scientists designed and tested peptides that resemble ACE2 enough to convince the coronavirus to bind to them, an action that blocks the virus’s ability to actually get inside the cell. “Our goal is that any time SARS-CoV-2 comes into contact with the peptides, the virus will be inactivated. This is because the virus Spike protein is already bound to something that it needs to use in order to bind to the cell,” said Amit Sharma, co-lead author of the study and assistant professor of veterinary biosciences at Ohio State. “To do this, we have to get to the virus while it’s still outside the cell.”
The study is published in the January issue of the journal Bioconjugate Chemistry.
The Ohio State team envisions delivering these manufactured peptides in a nasal spray or aerosol surface disinfectant, among other applications, to block circulating SARS-CoV-2 access points with an agent that prevents their entry into target cells. “With the results we generated with these peptides, we are well-positioned to move into product-development steps,” said Ross Larue, co-lead author and research assistant professor of pharmaceutics and pharmacology at Ohio State.
SARS-CoV-2, like all other viruses, requires access to living cells to do its damage – viruses hijack cell functions to make copies of themselves and cause infection. Very rapid virus replication can overwhelm the host system before immune cells can muster an effective defense. One reason this coronavirus is so infectious is because it binds very tightly to the ACE2 receptor, which is abundant on cells in humans and some other species. The Spike protein on the SARS-CoV-2 surface that has become its most recognizable characteristic is also fundamental to its success in attaching to ACE2. Recent advances in crystallizing proteins and microscopy have made it possible to create computer images of specific protein structures alone or in combination – such as when they bind to each other.
With these peptides, we have identified the minimal contacts needed to inactivate the virus
Amit Sharma
Sharma and his colleagues closely examined images of the SARS-CoV-2 Spike protein and ACE2, zooming in on precisely how their interactions occur and what connections are required for the two proteins to lock into place. They took notice of a spiral ribbon-like tail on ACE2 as the focal point of the attachment, which became the starting point for designing peptides. “Most of the peptides we designed are based on the ribbon contacting the Spike,” said Sharma, who also holds a faculty appointment in microbial infection and immunity. “We focused on creating the shortest possible peptides with the minimum essential contacts.”
The team tested several peptides as “competitive inhibitors” that could not only securely bind with SARS-CoV-2 Spike proteins, but also prevent or lower viral replication in cell cultures. Two peptides, one with the minimum contact points and another larger one, were effective at reducing SARS-CoV-2 infection in cell studies compared to controls. Sharma described these findings as the beginning of a product-development process that will be continued by the team of virologists and pharmaceutical chemists collaborating on this work. “We are taking a multipronged approach,” Sharma said. “With these peptides, we have identified the minimal contacts needed to inactivate the virus. Going forward we plan to focus on developing aspects of this technology for therapeutic purposes. The goal is to neutralize the virus effectively and potently, and now, because of the emergence of variants, we’re interested in assessing our technology against the emerging mutations.”
Source: Ohio State University
23.02.2021