News • Bioorganic chemistry
Why M. tuberculosis is so resistant to drugs and immune defenses
A consortium of researchers from Russia, Belarus, Japan, Germany and France led by the Skolkovo Institute of Science and Technology have uncovered the way in which Mycobacterium tuberculosis survives in iron-deficient conditions by utilizing rubredoxin B, a protein from a rubredoxin family that play an important role in adaptation to changing environmental conditions.
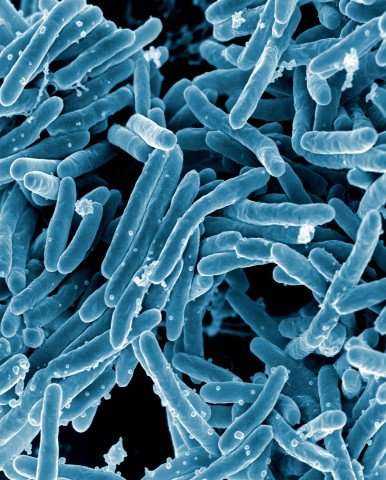
Image source: NIAID, Mycobacterium tuberculosis Bacteria (16843981465), CC BY 2.0
The new study is part of an effort to study the role of M. tuberculosis enzymes in developing resistance to the human immune system and medication. The paper was published in the journal Bioorganic Chemistry.
According to the World Health Organization, every year 10 million people fall ill with tuberculosis and about 1.5 million die from it, making it the world’s top infectious killer. The bacterium that causes TB, Mycobacterium tuberculosis, is notorious for its ability to survive within macrophages, cells of the immune system that destroy harmful bacteria. Continuing spread of drug resistance of M. tuberculosis to widely used therapeutics over recent decades became a substantial clinical problem. In this regard, the identification of novel molecular drug targets and deciphering the molecular mechanisms of drug resistance are of pivotal importance.
Natallia Strushkevich, Assistant Professor at the Skoltech Center for Computational and Data-Intensive Science and Engineering (CDISE), and her colleagues studied the crystal structure and function of rubredoxin B (RubB), a metalloprotein that ensures the proper functioning of cytochrome P450 (CYP) proteins essential to bacterial survival and pathogenicity. The team hypothesizes that M. tuberculosis switched over to more iron-efficient RubB to survive iron starvation when granulomas are formed (these are largely unsuccessful attempts at defense against TB by the immune system).
Our findings indicate that M. tuberculosis has its own xenobiotics transformation system resembling human drug metabolizing system
Natallia Strushkevich
“During the long-term co-evolution with mammals, M. tuberculosis developed a variety of strategies to subvert or evade the host innate immune response, from recognition of the bacterium and phagosomal defenses within infected macrophages, to adaptive immune responses by antigen presenting cells. Iron assimilation, storage and utilization is essential for M. tuberculosis pathogenesis and also involved in emergence of multi- and extensively-drug resistant strains. Heme is the preferable iron source for M. tuberculosis and serves as a cofactor for various metabolic enzymes. Based on our finding, we linked rubredoxin B to heme monoooxygenases important for metabolism of host immune oxysterols and anti tubercular drugs. Our findings indicate that M. tuberculosis has its own xenobiotics transformation system resembling human drug metabolizing system,” explains Natallia Strushkevich.
The expert elaborates: New targets for drug design efforts are in great demand and the cytochrome P450 enzymes have emerged as novel targets for the development of tuberculosis therapeutic agents. The classic approaches to block these enzymes are not straightforward. Finding the alternative redox partner, such as RubB, enables further understanding of their function in different host microenvironments. This knowledge could be exploited to identify new ways to block their function in M. tuberculosis. Earlier research by the consortium showed that one of the CYPs enabled by RubB can act against SQ109, a promising drug candidate against multidrug-resistant tuberculosis. Another study focused on how Mycobacterium tuberculosis protects itself by intercepting human immune signaling molecules — a hurdle that limits drug discovery.
Source: Skolkovo Institute of Science and Technology
10.04.2021