Article • Surface-enhanced Raman spectroscopy
The lab-on-a-chip SERS platform
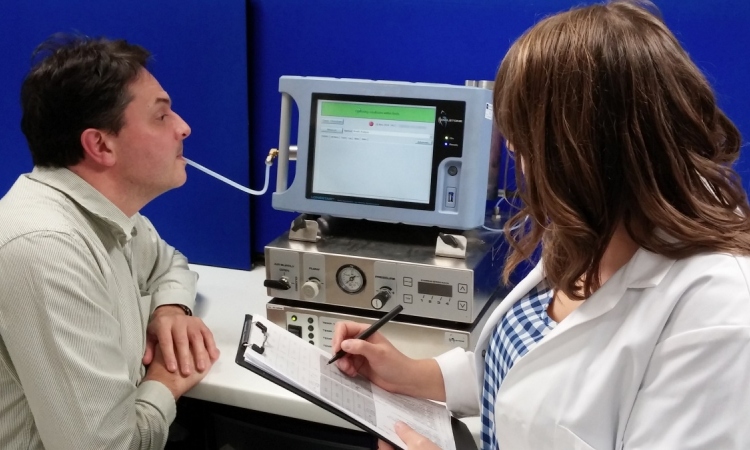
Analytically sensitive and specific detection of pharmaceuticals or metabolites in bodily fluids, as well as fast and reliable detection of human pathogens, are major challenges for instrument-based analytics in medical diagnostics.
Author's affiliations
1. Leibniz Institute of Photonic Technology (Leibniz IPHT); 2. Friedrich Schiller University Jena, Institute of Physical Chemistry and Abbe Centre of Photonics; 3. InfectoGnostics Science Campus; 4. Jena University Hospital (UKJ); 5. Klinikum rechts der Isar der TU München, Institute for Clinical Chemistry and Pathobiochemistry
Over the past few years the combination of surface-enhanced Raman spectroscopy (SERS) and microfluidic devices (Lab-on-a-Chip) has emerged as a perfectly suited technology to ensure an automated and reproducible measuring environment and/or to analyse nanolitre volumes.1
The key feature of SERS is its ability to detect with molecular-specificity minute concentrations of pharmaceutical substances. The required sensitivity is achieved by using plasmonic active metallic nanostructures, such as colloidal silver or gold nanoparticles, or nano-structured or micro-structured metallic surfaces. Furthermore, this method can be applied directly, i.e. without complex sample purification protocols, to detect individual pharmaceutical agents or metabolites in complex human matrices, such as urine, plasma or sputum, in concentrations of few mg/mL and even below µg/mL.
While SERS offers enormous potential in bioanalytics, it is not yet part of the routine diagnostic workup. This might to some extent be due to the high quality requirements with regard to SERS-active nanostructures which have to ensure reproducible signal intensity. Microfluidic devices, i.e. Lab-on-a-Chip can overcome the drawbacks of SERS such as the hard-to-control aggregation of the colloidal metal nanoparticles.
Currently, there are two types of Lab-on-a-Chip SERS platforms: flow-through or continuous flow platforms and segmented or droplet-based platforms.1 In flow-through platforms either metallic colloidal nanoparticles are mixed with the samples to be analysed, or stationary metallic nanostructures are integrated in the channel system. This approach avoids the evaporation of liquids, which occurs during conventional SERS measurements on open platforms. A disadvantage, however, is that sample liquids permanently wet the channel walls, which may result in the so-called memory effect (Memory effect describes a situation when the molecules of the sample get enriched over time on the surface of the channels. This may result in an overestimation of the detection limits).
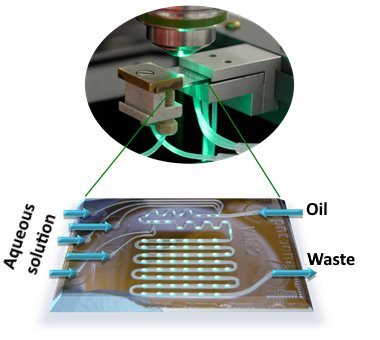
On a segmented flow platform, particularly droplet-based SERS (Figure 1), cross-contamination can be fully avoided, since aqueous droplets are transported in an oily matrix. Thus there is no contact between the sample, resp. the metallic colloids and the channel walls. Furthermore, each droplet can be considered a micro-cuvette or a micro-reactor. Thus many spectra can be fed quickly and in an automated way into a database.
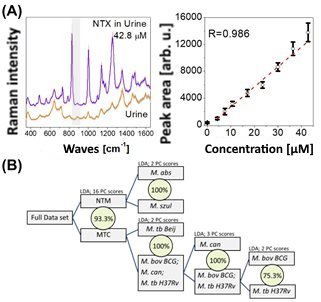
The authors’ working group successfully used the droplet-based Lab-on-a-Chip SERS technology to detect pharmaceuticals and metabolites in bodily fluids and to identify mycobacteria. In these studies human urine was spiked with the antibiotic levofloxacin. Concentrations between 0.45 mM (162.6 μg/mL) and 1.8 mM (650.5 μg/mL) were detected.2 According to relevant literature the expected levofloxacin concentrations in urine are 1.38 mM ± 0.68 mM and a measured minimum concentration of 0.45 mM 4 hours post administration of 500 mg. Furthermore, the standard addition method was successfully used to detect nitroxoline (NTX) in human urine.3 Figure 2A shows a typical SERS spectrum of NTX and the concentration-dependent evolution of Raman-intensity of a marker band. The results presented above show that Lab-on-a-Chip SERS is a promising bioanalytical tool for urine analysis.
Moreover, enzyme activity of thiopurine methyltransferase (TPMT) in lysed erythrocytes was successfully determined.4 This involved tracking the transformation of the drug 6-mercaptopurine into 6-methylmercaptopurine, which in turn allowed conclusions regarding TPMT activity. No and normal TPMT activity were distinguished with 96% accuracy.
As far as the detection of pathogenic bacteria is concerned, the Pseudomonas aeruginosa-specific metabolite pyocyanine can be quantified with SERS.5 Using a robust SERS substrate pyocyanine was detected in artificial sputum depending on the concentration in the relevant area in cystic fibrosis patients who frequently show Pseudomonas aeruginosa colonisation.6
Finally, Lab-on-a-Chip SERS was successfully used to identify mycobacteria in order to facilitate physicians’ decision-making regarding medication.7 Figure 2B shows the chemometric model for the identification of bacteria.
To summarise: Lab-on-a-Chip SERS technology has high potential for clinical applications. Further studies with clinical samples are required to confirm these initial findings.
References:
06.08.2019