Deep Learning
Philips and LabPON plan to create world’s largest pathology database
Royal Philips (NYSE: PHG, AEX: PHIA) and LabPON, the first clinical laboratory to transition to 100% histopathology digital diagnosis, today announced its plans to create a digital database of massive aggregated sets of annotated pathology images and big data utilizing Philips IntelliSite Pathology Solution1. The database will provide pathologists with a wealth of clinical information for the development of image analytics algorithms for computational pathology and pathology education, while promoting research and discovery to develop new insights for disease assessment, including cancer.
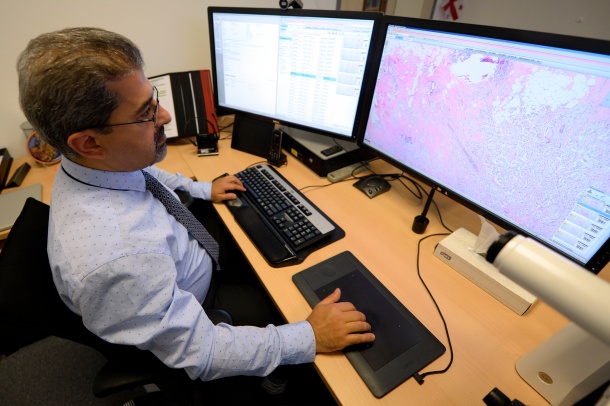

Deep learning algorithms have the potential to improve the objectivity and efficiency in tumor tissue diagnosis. In recent years, ‘deep learning’ techniques for image analysis have quickly become the state of the art in computer vision and has surpassed human performance in a number of tasks2. The challenge for executing deep learning techniques is having access to a database with sufficient high volume and high quality data from which to develop the algorithms. As one of the largest pathology laboratories in the Netherlands, LabPON will contribute its repository of approximately 300,000 whole slide images (WSI) they prospectively create each year to the database. This will contain de-identified datasets of annotated cases that are manually commented by the pathologist, and will comprise of a wide variety of tissue and disease types, as well as other pertinent diagnostic information to facilitate deep learning.
“Deep learning focuses on the development of advanced computer programs that automatically understand and digitally map tissue images in considerable detail: The more data available, the more refined the computer analysis will be.” Said Peter Hamilton, Group Leader Image Analytics at Philips Digital Pathology Solutions. “Together, LabPON and Philips have the competence and skills to realize this.”
During a time where the pathologist shortage is mounting and cancer caseloads are increasing3,4, the accurate diagnosis and grading of cancer has become increasingly complex, placing significant pressures on pathology services. Technologies such as computational pathology, could help pathologists with tools to work in the most efficient way possible.
“The role of the pathologist remains important by making the definitive diagnosis, which has a high impact on the patient’s treatment. Software tools could help to relief part of the pathologists’ work such as identifying tumor cells, counting mitotic cells or identifying perineural and vaso-invasive growth, as well carrying out measurements in a more accurate and precise way,” said Alexi Baidoshvili, pathologist at LabPON. “This ultimately could help to improve the quality of diagnosis and make it more objective.”
Next to the development of computational algorithms for diagnostic use, Philips intends to make available the database to research institutions and other partners through its translational research platform. This could enable selected parties to interrogate and combine massive datasets with the goal to discover new insights that ultimately could be translated into new personalized treatment options for patients.
Philips is showcasing its portfolio of pathology solutions in booth #202 at the The United States and Canadian Academy of Pathology (USCAP) 2017 Annual Meeting. For more information on Philips’ presence at USCAP, visit www.philips.com/digitalpathology and follow @Philips_Path for #USCAP17 updates throughout the event.
1 Philips IntelliSite Pathology Solution is CE-IVD marked for use in primary diagnosis. In the United States, the Philips IntelliSite Pathology Solution pending review of a request for de novo classification.
2 Kaiming He Xiangyu et al. Delving Deep into Rectifiers: Surpassing Human-Level Performance on ImageNet Classification. And
LeCun, Yann, Yoshua Bengio, and Geoffrey Hinton. “Deep learning.” Nature 521, no. 7553 (2015): 436-444
3 The Royal College of Pathologists, https://www.rcpath.org/profession/workforce/workforce-planning.html, Accessed December 2016.,
4 International Agency for Research on Cancer and Cancer Research UK. World Cancer Factsheet. Cancer Research UK, London, 2014.
Source: press release, Carver Wilde Communications
09.03.2017