News • Proof of concept
Surveillance system tracks Covid infection hotspots in hospital
A University of Manchester team has applied new techniques to detect and track the transmission of Covid-19 in hospital.

Image source: Unsplash/United Nations Covid-19 Response / created by Russell Tate. Submitted for United Nations Global Call Out To Creatives
The proof of concept system combines the movement and interaction of staff and patients with genomic sequencing of the virus, helping to signpost how best to improve patient pathways, staff movement and reduce risk. They identified hotspots within hospitals where patients and staff had likely been in contact and shared similar or identical variants of the virus SARS-CoV-2- which causes Covid-19. The data collected between March and June 2020, was the result of a unique collaboration between Manchester scientists, clinical staff and hospital executives whose results are published in the journal Elife.
The work, which took place across five hospitals in North West England, could have an important impact on infection control. The team applied genome sequencing to throat and nasal swabs obtained from 173 healthcare workers and patients. Viruses, like all organisms, accumulate small genetic changes through time which are usually completely neutral and don’t give the virus any additional advantages. The Manchester team took advantage of the natural genetic changes in the virus to understand how closely related samples from different staff and patients affected by the virus were. If the Covid-19 genome from one affected individual appears almost identical to that of another, this shows they share a common, nearby source of infection. Combining this information with data on patient and staff movement can identify where and when clusters of infection occur.
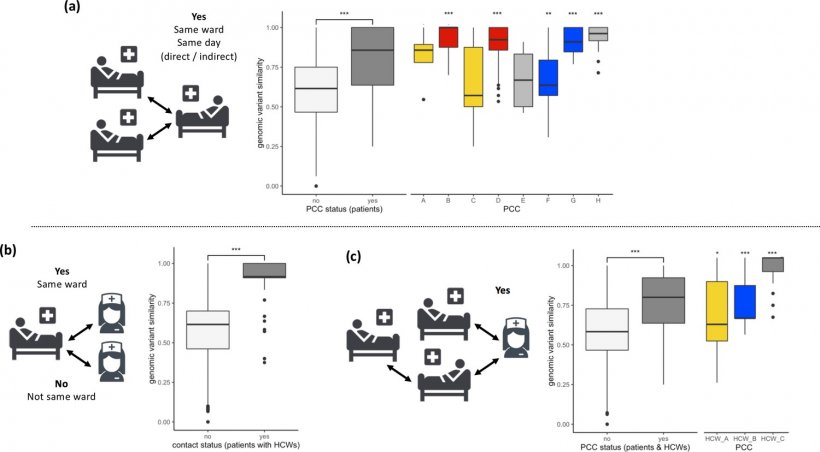
Image source: Ellingford et al., Elife 2021 (CC BY 4.0)
While Covid-19 variants are identified using genomic sequencing in a process that can take at least a week to complete, the team are confident the process can now be carried out as quickly as 48 hours. The approach is therefore in a position to be scaled up at pace. Dr Jamie Ellingford from The University of Manchester said: “The methods applied in this study to completely characterise what the virus looks like in each sample goes above and beyond routine testing strategies. And that can enable identification of areas of hospitals where outbreaks are occurring and help alert infection control teams.”
An important tool in the infection control armoury is viral genome sequencing, which offers a realistic possibility to track and identify root-causes of hospital-acquired transmissions
Jamie Ellingford
A potential limitation to the approach is that at any given time point, two individuals can share a similar variant by chance, rather than as a result of interrelated infections. To overcome that, the team sequenced the virus in people collected from the Emergency Department and across over 30 hospital sites. Using data from this wider group collected at a similar time point, they could identify clusters of individuals who shared viruses more genetically similar than would be expected by chance. Dr Ellingford said: “It is extremely important to understand the effectiveness of infection control methods if we are to reduce and prevent SARS-CoV-2 transmission in hospital. While vaccines may reduce risk to individuals in hospital, the risk of infection will still exist and infection control is strongly needed to ensure patient and staff safety. We think, however, an important tool in the infection control armoury is viral genome sequencing, which offers a realistic possibility to track and identify root-causes of hospital-acquired transmissions. It is able to alert us to individuals who have been in contact during a given period and share genetically similar viral samples. And that can lead to targeted interventions and ultimately prevent avoidable harm to vulnerable individuals who acquire Covid-19 in hospital.”
Graeme Black is Professor of Genetics and Ophthalmology at the University of Manchester. He said “Once hospitals have identified clusters there are a range of measures they can take to make them safe. With this information hospital managers can, for example, evaluate existing infection control practices and easily check which ones are working most effectively. The use of PPE could also be adapted according to where a cluster might be. But we also suggest these data support the widespread adoption of screening strategies for healthcare workers who may be presymptomatic or asymptomatic shedders of SARS-CoV-2 who are important contributors to SARS-CoV-2 outbreaks. We also think that developing methods to more accurately track movement could be useful in hospital to extend the characterisation of contacts between individuals and to understand the accuracy of the assumptions enforced in this study.”
Source: University of Manchester
01.04.2021





