Image source: Goethe University Frankfurt
News • Coronavirus genome folding
Researchers prepare for “SARS-CoV-3”
For the first time, an international research alliance has observed the RNA folding structures of the SARS-CoV2 genome with which the virus controls the infection process.
Since these structures are very similar among various beta corona viruses, the scientists not only laid the foundation for the targeted development of novel drugs for treating Covid-19, but also for occurrences of infection with new corona viruses that may develop in the future.
The researchers present their findings in the journal Nucleic Acids Research.
We hope that we can contribute to being better prepared for future ‘SARS-CoV3’ viruses
Harald Schwalbe
The genetic code of the SARS-CoV2 virus is exactly 29,902 characters long, strung through a long RNA molecule. It contains the information for the production of 27 proteins. This is not much compared to the possible 40,000 kinds of protein that a human cell can produce. Viruses, however, use the metabolic processes of their host cells to multiply. Crucial to this strategy is that viruses can precisely control the synthesis of their own proteins.
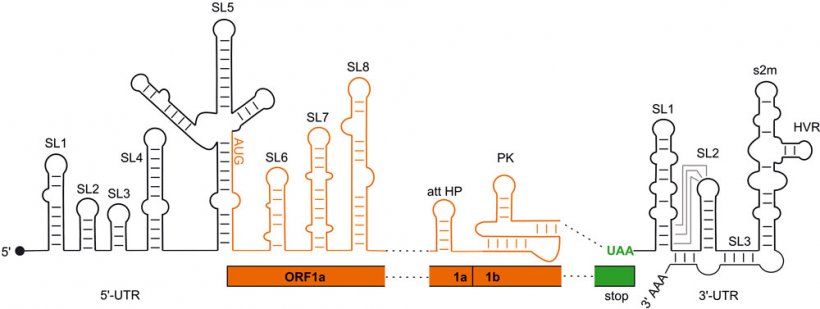
SARS-CoV2 uses the spatial folding of its RNA hereditary molecule as control element for the production of proteins: predominantly in areas that do not code for the viral proteins, RNA single strands adopt structures with RNA double strand sections and loops. However, until now the only models of these foldings have been based on computer analyses and indirect experimental evidence.
Now, an international team of scientists led by chemists and biochemists at Goethe University and TU Darmstadt have experimentally tested the models for the first time. Researchers from the Israeli Weizmann Institute of Science, the Swedish Karolinska Institute and the Catholic University of Valencia were also involved.
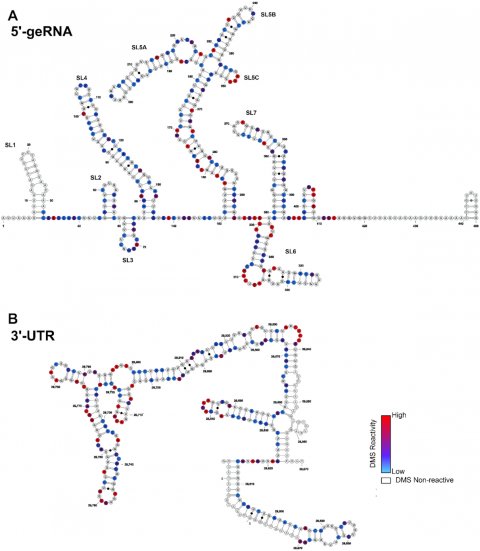
Image source: Wacker et al., Nucleic Acids Research 2020 (CC BY-NC 4.0)
The researchers were able to characterise the structure of a total of 15 of these regulatory elements. To do so, they used nuclear magnetic resonance (NMR) spectroscopy in which the atoms of the RNA are exposed to a strong magnetic field, and thereby reveal something about their spatial arrangement. They compared the findings from this method with the findings from a chemical process (dimethyl sulphate footprint) which allows RNA single strand regions to be distinguished from RNA double strand regions.
The coordinator of the consortium, Professor Harald Schwalbe from the Center for Biomolecular Magnetic Resonance at Goethe University Frankfurt, explains: “Our findings have laid a broad foundation for future understanding of how exactly SARS-CoV2 controls the infection process. Scientifically, this was a huge, very labour-intensive effort which we were only able to accomplish because of the extraordinary commitment of the teams here in Frankfurt and Darmstadt together with our partners in the Covid-19-NMR consortium. But the work goes on: together with our partners, we are currently investigating which viral proteins and which proteins of the human host cells interact with the folded regulatory regions of the RNA, and whether this may result in therapeutic approaches.”
Worldwide, over 40 working groups with 200 scientists are conducting research within the Covid-19-NMR consortium, including 45 doctoral and postdoctoral students in Frankfurt working in two shifts per day, seven days of the week since the end of March 2020.
Schwalbe is convinced that the potential for discovery goes beyond new therapeutic options for infections with SARS-CoV2: “The control regions of viral RNA whose structure we examined are, for example, almost identical for SARS-CoV and also very similar for other beta-coronaviruses. For this reason, we hope that we can contribute to being better prepared for future ‘SARS-CoV3’ viruses.”
Source: Goethe University Frankfurt
23.11.2020