News • Direct from whole blood samples
LiDia demonstrates rapid and sensitive BSI identification
DNAe, the inventor of semiconductor-based genomic analysis technologies, and the developer of a new, game-changing test for bloodstream infections that can lead to sepsis announced new data on its test for bloodstream infections, LiDia BSI.
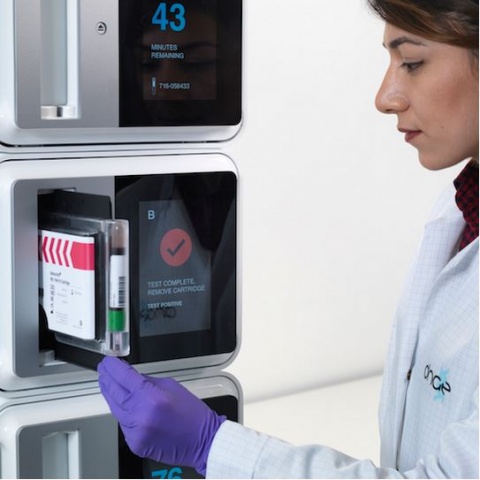

The data demonstrates the ability of the LiDia BSI closed cartridge-based test to rapidly identify low levels of bacterial and fungal pathogens and resistance markers direct from whole blood, including an example of successful automation. The data were presented as a poster (ID31) at the Association for Molecular Pathology (AMP) Annual Meeting 2017 in Salt Lake City, UT, USA (16th – 18th November). LiDia BSI was used to accurately detect the most common pathogens linked to serious bloodstream infections, including the superbug methicillin resistant S. aureus (MRSA), less frequent pathogens that are often mistreated empirically, and important resistance markers. The authors of the poster at DNAe confirmed end-to-end functionality of the LiDia BSI workflow, correctly identifying pathogens and resistance genes at 1 colony-forming unit per ml (CFU/ml). Total turnaround time was less than 5 hours, with current automation of the process expected to reduce the total process time down to less than 3 hours.
The poster presentation includes data on the automation of the LiDia BSI workflow, used to correctly detect the pathogen, K. pneumoniae at 2.5 CFU/ml. Currently, the standard of care for diagnosis of bloodstream infection is via blood culture, which can take two to six days to produce results. For serious bloodstream infections, which can lead to the life-threatening condition of sepsis, early diagnosis and administration of antimicrobials targeting the causative pathogen is the single most important factor in reducing mortality and morbidity. Reducing time to diagnosis from days to hours therefore has potentially lifesaving implications. On track for CE marking in 2018, LiDia BSI will reduce unnecessary prescription of antibiotics by providing actionable results directing targeted treatment in just a few hours, direct from a raw blood specimen.
Detection of pathogens in healthy donor blood samples spiked at 1 CFU/ml
LiDia BSI correctly identified the following pathogens in healthy blood samples individually spiked at 1 CFU/ml: A. baumannii, K. oxytoca, K. pneumoniae, P. mirabilis, E. faecalis, E. faecium, S. aureus, C. albicans, and C. glabrata. The absence of resistance markers in these samples was correctly confirmed in K. pneumoniae, E. faecalis, E. faecium, and S. aureus. In addition, the presence of both pathogen and resistance marker was confirmed for contrived whole blood specimens with vancomycin-resistant E. faecalis, vancomycin-resistant E. faecium, MRSA, and K. pneumoniae harbouring blaKPC – all targets spiked at 1 CFU/ml.
Detection of pathogens in clinical whole blood samples by LiDia BSI
The ability to generate robust data from blood within hours, compared to days, will be transformational for patients with bloodstream infection
Steve Allen
Clinical specimens of MRSA and methicillin-sensitive S. aureus were correctly identified directly from whole blood via detection of S. aureus and the presence or absence of mecA/C, concordant with blood culture results of the samples. The abstract, published in The Journal of Molecular Diagnostics, vol 19(6), pp. 980, is available online here: http://jmd.amjpathol.org/article/S1525-1578(17)30482-8/pdf
The full data are available here: http://dnae.com/assets/amp-poster-2017-final-online.pdf
David Davidson, Chief Scientific Officer at DNAe and author on the poster said, “It’s exciting to be able to clearly demonstrate the ability of LiDia BSI to detect pathogens and antimicrobial resistance present even at low levels, directly from raw blood samples. We continue to process clinical samples with some representative data points presented at this year’s AMP.”
Professor Chris Toumazou, Founder and Executive Chairman at DNAe and author on the poster said, “We are pleased to share this outstanding dataset including an automated exemplar at low limits of detection. Through the eventual automation of the entire workflow, LiDia BSI has huge potential to cut down time to diagnosis for patients with bloodstream infection. We look forward to sharing larger datasets from our ongoing clinical testing program.”
Dr Steve Allen, CEO of DNAe Group Holdings, commented: “These data provide evidence of the sensitivity, breadth of pathogen coverage, inclusion of resistance genes, and speed of the LiDia platform and test offering. Presenting these data at the AMP 2017 Annual Meeting is an important stepping stone as we move swiftly towards commercialization of the LiDia BSI test. The ability to generate robust data from blood within hours, compared to days, will be transformational for patients with bloodstream infection. The next stage in the development process will be to fully integrate the workflow into the cartridge-based test to allow the system to deliver clinically actionable results in less than 3 hours.”
Source: DNAe
20.11.2017