Article • Wishlist
PACS and imaging biobank assets combined
Report: Marcel Rasch
ECR 2016 - Don't miss
Friday, March 4, 08:30–10:00, Studio 2016
Biobanks meet imaging

Two problems have arisen. The first is the storage of research image data; the second is the extraction of imaging biomarkers. Radiologists may wonder whether the hospital PACS can solve these problems. Working with a PACS to provide better patient care by analysing their images and obtaining reports to improve diagnoses is nothing new to most clinicians. PACS provides storage and access to all images acquired during the diagnostic and therapeutic phase of the disease. Nowadays it can be integrated in the digital electronic patient record (EPR). For radiologists, a PACS provides a workspace for planning and reporting image exams. More importantly, the integration of image analysis tools allows the creation of 2-D or 3-D reconstructions and the extraction of imaging biomarkers. It is foreseen that structured reporting will also rely on the digital environment of the PACS and integrated thin client analysis tools.
However, storage of image data for biobanking has specific requirements and there are major differences between a PACS and an image biobank. A biobank contains anonymised data and is accessible by researchers that have permission from the principal investigators of the biobank. PACS data contains the name of the patient and are accessible by all physicians involved in the treatment of this patient. Image storage in most research projects is not stable and robust, future data access is not secured and a secure data access is not always maintained. In contrast, a PACS is very stable and robust, data can be stored for more than 20 years, a secure access is guaranteed and loss of data is minimized.
An image biobank needs a DICOM viewer and additional analysis tools to review the image data and to perform analysis and interpretation of these data by humans. One of the main advantages of PACS is the presence of these tools. ‘In the clinical environment we have a very nice PACS in which to analyse these data, but what we do not have is the possibility of using these analytical tools from the PACS in a research environment. Making full use of the options provided by a PACS system in a research infrastructure could take science to a completely new level,’ he adds.
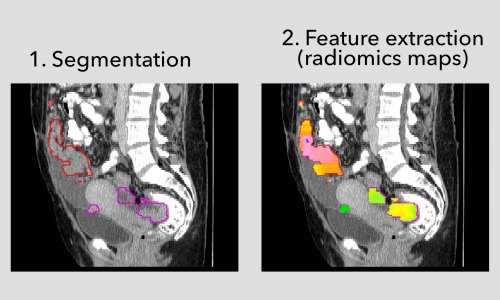
An image biobank with a well-defined structure allows query and retrieve of image data based on available metadata. In addition, multiple imaging biomarkers can be extracted by fully automated image analysis tools with integrated pipelines that make use of GRID computing. Images of hundreds of patients can be registered, relevant structures can be segmented and quantitative biomarkers can be extracted and automatically stored in the database. A PACS does not allow query and retrieve on a large scale, nor the fully automated analysis of hundreds of image data sets. ‘What we need from a research perspective is just not available in a PACS, although some PACS features would be very useful for research.’In the clinical environment, the assessment of image information relies on a description that is provided by an imaging expert: the radiologist. In a research setting intelligent machine learning tools increasingly let the systems learn step by step how to read and interpret the images, but this requires that image data are prepared sufficiently and are made comparable and analysable by computers.
Finally, in biobanking image biomarkers can be related to data in the other -omics domains. However, these tools are not available as yet in the clinical setting.
Closing the gap
‘Currently we have a mismatch that must be solved,’ he states, specifying his demand: ‘We should try to close the gap between the research infrastructure and the PACS system. In other words, using the tools that are available in a PACS in a research environment and using the PACS for more advanced research analysis would be the optimum. Combining the advantages of both approaches would be the best solution.’ This includes a strategy of storage of anonymised data in a split PACS system. One for clinical use and one for research use with a separate access to the two domains, and implementation of automated algorithms that can make use of multiple image datasets that, for example, can detect lesions and automatically measures their size. ‘There have to be advanced algorithms that can analyse large scale data and export the quantitative imaging biomarkers to one’s research database,’ Van der Lugt believes. ‘This approach allows the use of the data richness of the images that are available for most of the patients included in biobanks.
Profile:
Aad van der Lugt, Professor in Neuroradiology and Head-Neck-Radiology at Erasmus MC, Rotterdam, Netherlands is director of the neuroradiological research programme. In 2007, he expanded his research into imaging biomarkers in large population-based studies. Within these epidemiology studies, the professor is responsible for the imaging infrastructure. He is one of the cofounders of EPI2 (European Population Imaging Infrastructure), co-applicant of NL-BBMRI 2.0 (Biobanking and Biomolecular Resources Research Infrastructure in the Netherlands) and responsible for Population Imaging in the Euro-BioImaging (ESFRI) project. He is also a member of the research committee of the European Society of Radiology.
03.03.2016