News • RNA sequencing
Overcoming a major barrier to developing liquid biopsies
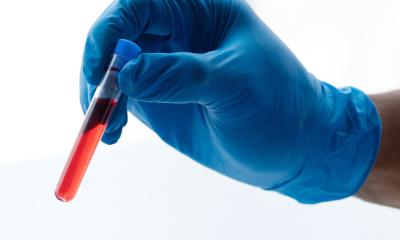
The idea of testing blood or urine to find markers that help diagnose or treat disease holds great promise. A new study helps set standards for RNA sequencing, improving consistency and laying the foundation to move the field forward.

The idea of testing blood or urine to find markers that help diagnose or treat disease holds great promise. But as technology has improved to allow researchers to examine tiny fragments of RNA, one major problem has led to limited success. “Different people are using different methods to sequence small RNA, and sometimes getting different results. If it keeps going on like that, it will be hard for the field to make progress,” says Muneesh Tewari, M.D., Ph.D., professor of internal medicine and biomedical engineering at Michigan Medicine.
Tewari’s lab led a group of nine labs across the United States and the Netherlands, brought together through the National Institutes of Health, that sought to solve this problem. The consortium tested nine different methods for RNA sequencing to understand and standardize the best way to sequence small RNAs. The goal was to create a process that could be reproduced from one lab to the next.
A liquid biopsy relies largely on the ability to sequence small RNA such as microRNA. These tiny cellular fragments can become altered in diseases such as cancer, providing a clue to help spot disease in its earliest stages. But blood or urine contain only a tiny amount of RNA outside of cells, making it challenging to sequence. “Liquid biopsy for RNA is an exciting new field for diagnostics. But the field needed this kind of consortium to come together, because of the challenge of different methods leading to results that are not reproducible,” Tewari says. He also notes a strength of this work is in bringing together varied expertise, from molecular biologists to computational and bioinformatics specialists.

Credit: Sigrid Knemeyer, SciStories LLC
Standard procedures necessary
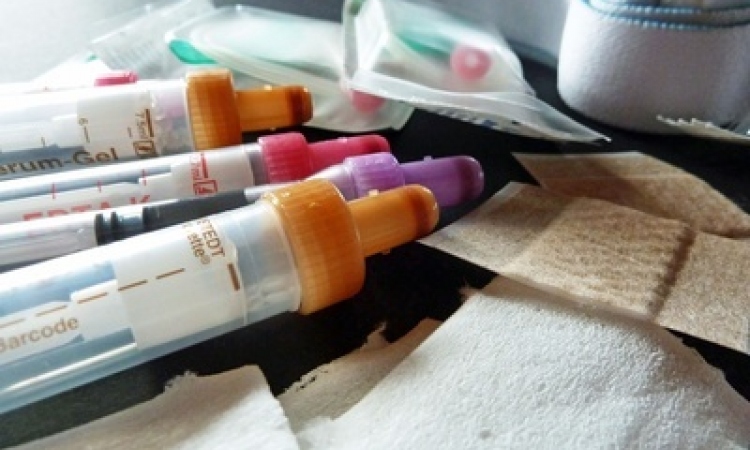
For this study researchers prepared samples identically and sent them across the country for each of the nine labs to analyze. Each lab used multiple testing protocols to sequence four different samples, including a plasma sample and three synthetic RNA samples. Altogether, they tested nine different sequencing protocols, including four commercially available kits and five protocols developed by the labs. The combined data yielded more than 5 billion sequencing reads.
“We realized that not only different methods produce different results, but also any small change within a given protocol can introduce an important degree of variation. In order to compare results across labs, it is key to use a common and highly standardized protocol,” says lead study author Maria D. Giraldez, M.D., Ph.D., a post-doctoral research fellow in Tewari’s lab.
Researchers found that different methods used for sequencing produced different, often inaccurate, estimates of how abundant any individual marker was. The methods developed by the consortium labs improved the accuracy of these estimates. When RNA sequencing was used to compare the relative amounts of individual microRNAs between different samples, however, all the methods produced accurate and reproducible estimates.
“We found there was not a lot of variability if you used the same protocol across multiple labs,” says study author Ryan Spengler, Ph.D., a post-doctoral fellow in Tewari’s lab. “This means, if you want to coordinate a study between different labs, the key is to keep to the same protocol – whatever it is. Then you can compare your results.”
The analysis lays a foundation to help researchers create standard procedures around their protocols, and positions the field of RNA sequencing and liquid biopsies to move forward. The researchers have made the synthetic reference material available, which means researchers across the country can run their test and compare results to what the consortium of labs found.
Source: Michigan Medicine - University of Michigan
17.07.2018