Article • A more integrative approach to digital pathology
imCMS: The door to simple, cheap, reliable bio-stratification
Bringing molecular and digital pathology closer together through a more integrative approach can lead to clear advantages for diagnostic and research workflows.
Report: Mark Nicholls
During the recent Digital Pathology and AI Congress (London) and in his keynote presentation ‘Molecular and digital pathology - the value of an integrative approach’, Professor Viktor Koelzer explored the benefits and paid particular attention to colorectal cancer (CRC) during the event. ‘It’s an exciting time in pathology as we better connect tissue morphology and molecular changes using digital pathology and artificial intelligence,’ he told delegates.

Koelzer, who is Attending Pathologist and Assistant Professor at the Institute of Pathology and Molecular Pathology at University of Zurich and University Hospital Zurich, explored recent paradigm changes in clinical diagnostics and research workflows with the potential of predicting molecular features from image data. ‘Information from surgical pathology feeds into molecular testing, but we rarely make the loop back. Today we are starting to close this gap,’ said Koelzer.
Looking at tumour annotation for molecular analysis, he said challenging areas include the limited reproducibility of visual pathology review, inconsistency with bioinformatic prediction of tumour cell content using sequencing data and the resulting limited correlation of standard pathology assessment with DNA and RNA yield from clinical samples. As the ability to detect somatic variants in cancer samples drives personalised therapy, better tissue classification strategies could help to improve clinical diagnostic workflows.
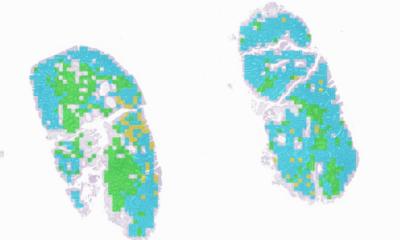
‘An area of development is tissue segmentation by supervised machine learning for the estimation of tumour cell percentage by digital pathology,’ he said. This approach allows correlation between different levels of information and provides accurate area information for each tissue component on a given histology slide.
Excellent reproducibility
‘It allows us to look at the cell-level composition of histology slides at great detail and with excellent reproducibility,” he added, “we can thereby gain additional information from pathology review in an automated fashion that would otherwise be very laborious to obtain.’
Indeed, initial data showed that tissue composition analysis with DNN (deep neural networks) allows analytical robustness, automatisation and standardisation and provides high reproducibility at single cell resolution. DNA-based tumour purity estimates are more accurate than visual view or deconvolution from genome-wide omic platforms which, he said, tend to under- as well as over-estimate tumour purity respectively. Therefore, digital pathology review using DNN could be used to inform downstream molecular analyses better and investigate tissue-based metrics as potential biomarkers in clinical trials.
CRC has significant potential and is a current area of focus for his research team at USZ in collaboration with Professor Jens Rittscher and Professor Tim Maughan at the University of Oxford. As part of the CRUK and MRC funded Stratification in Colorectal Cancer or S:CORT consortium, they have previously developed a cheaper, faster method of cancer classification using artificial intelligence to analyse high-resolution images of histological slides. This allows the subclassification of colorectal tumours into one of four distinct consensus molecular subtypes (CMS) and gives an indication of optimal treatment strategies.
As histological grading is a poor predictor of disease progression, and CMS cannot be distinguished without gene expression profiling, a clinical need results for better prognostic/predictive indicators for the disease. ‘Clinical stratification using molecular data requires sequencing of tissue samples, which is costly and requires the availability of the relevant infrastructure and bioinformatics resource,’ he added, explaining that image analysis can be a cost-effective solution to interrogate morphological features as a surrogate for genetic/molecular information.
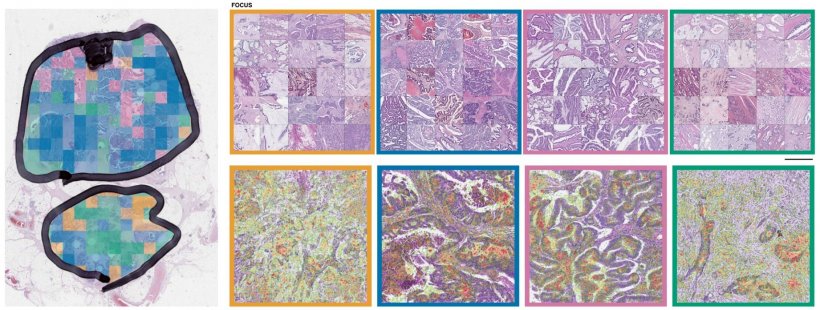
Biological understanding of CRS
‘We show that image-based CMS (imCMS) of CRC can assess clinical impact, produce a molecular profile and phenotype information for molecular subtypes of CRC.’ Using this information could improve our biological understanding of CRC potentially leading to better clinical stratification and treatment decisions, such as performing a surgical resection or giving adjuvant chemotherapy.
In conclusion, he said that Computational models can predict transcriptional subtype of CRC from standard histology sections. ‘imCMS makes sequencing information interpretable through association of morphology, molecular features and outcome data and classifies samples previously unclassifiable by RNA expression profiling. It also gives a novel insight into tumour heterogeneity, is highly prognostic; and classifies endoscopic biopsies and resection specimens of CRC enabling patient stratification in diverse clinical settings.’ Koelzer sees a promising outlook with a better integrative approach for molecular and digital pathology: ‘Molecular classifiers can be recognised and called from H&E images, opening the door to simple, cheap and reliable biological stratification within routine workflows and existing retrospective cohorts.’
Profile:
Viktor Koelzer is Assistant Professor at the Institute of Pathology and Molecular Pathology at University Hospital Zurich and Honorary Senior Clinical Researcher at the University of Oxford. He is passionate about technology in application to daily diagnostic practice and research, in particular improving patient care through the implementation of high-quality, science-driven computational image analysis approaches with a focus on gastrointestinal disease and tumour immunology.
07.05.2020