Article • TMAs in digital pathology
Infusing tissue microarrays with new, 'digital' life
The advent of digital pathology is helping to address some of the challenges surrounding tissue microarrays as they are integrated into the digital workflow, in some ways giving them ‘a new lease of life’, according to Professor Inti Zlobec, who spoke at the Digital Pathology and AI Congress in London last December.
Report: Mark Nicholls
As Head of the Translational Research Unit at the Institute of Pathology, University of Bern, she pointed out the challenges surrounding next-generation Tissue Microarray (ngTMA) and explored the benefits of coupling with digital pathology in translational research. A long work history with tissue microarrays indicates that digital pathology is now ‘vital to the whole process’ of how TMAs are constructed. Reflecting on the early days of TMAs from 1998 and the combination of different spots on one block, Zlobec said the idea uses less resources and material and, by adding associated pathological data, a research tool can be created that can last 5-10 years.

It was clear for some time that TMAs might be a valuable screening tool, and her team, which has a focus on colorectal cancer, was interested in tumour budding but the challenge was how to get budding on a TMA. ‘One solution was to incorporate digital pathology into the microarray workflow,’ she said. ‘The digital slide could be used to annotate exactly the region or multiple regions you wanted. By increasing the resolution, you could really see what you were putting on the TMAs.’
The next-generation Tissue Microarray approach, which utilises slide scanning and digital annotation as a basis for TMA construction, has been used in her lab since 2012. ‘The beauty of that is that you have fully annotated – and documented – spots and images. There was also the ability for every single spot to have clinical and pathological information and the clinical outcome for every single patient.’
Her Bern group created 720 TMA blocks with about 150,000 punches thumped physically into the blocks. While colorectal cancer was the main focus – about 30% of the blocks – the team also created tissue blocks for lung, pancreas, oesophagus, prostate and other areas.
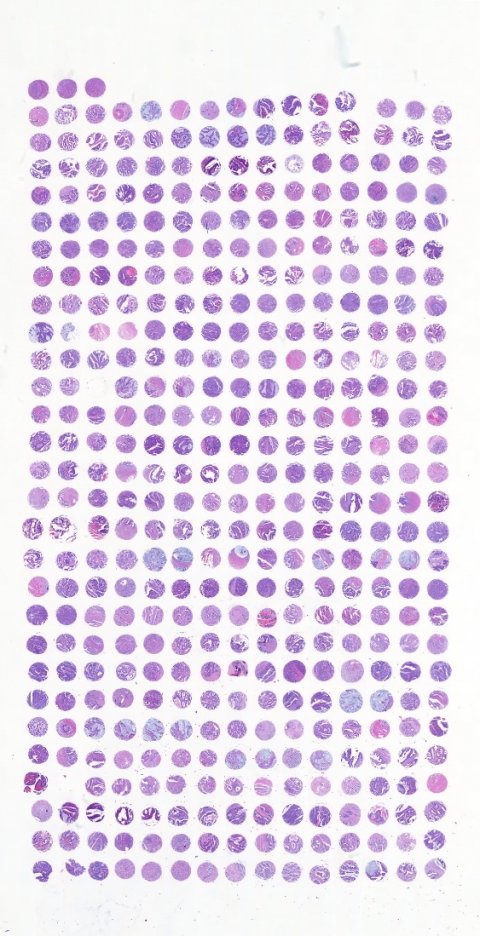

Ultimately the process led to more than one million images in the TMA archive from different tumour entities (after TMA cutting and staining) and all with the associated pathological annotation, histological information and clinical outcome… and potentially available for image analysis. ‘Now the problem is how to manage that number of TMA slides,’ she said. ‘A solution is with machine learning tools and having the TMA spots linked back to patients, whole slide images, annotative information and image analysis results, though this is still work in progress.’ Another question Zlobec posed was whether, with the ever-advancing technology, TMAs are still relevant and, she noted, the number of annual publications on the subject had plateaued in recent years at about 600. The original aim of TMAs of screening and resource efficiency, she said, was still relevant and, they can be used to study heterogeneity, biomarkers, and to capture the tumour environment. More recently, the Bern group has developed a tool (Scorenado) that provides an intermediate solution to image analysis, facilitating user-friendly visual assessment on digital slides. Today, TMAs are a resource in the machine learning era for classification or prognosis and prediction for patient outcomes and also for translation into clinics.
‘They also combine technology with molecular applications – we can take the punch we put on a TMA and then put it into a tube to do the sequencing, giving us the kind of documentation we really need,’ Zlobec explained, also pointing out that digital pathology also can complement mass spectrometry, identifying and validating new biomarkers. As digital pathology evolves, its added value to TMA use becomes clear. It improves, she pointed out, the quality of biomarker studies with precise ngTMAs that quickly capture ROIs; uses digital image analysis; improves documentation; aids biobanking and the accreditation process; it also offers high quality pathology standard annotation of images, and can be combined with molecular applications. Additionally, using the image as a biomarker adds a huge wealth of morphology information.
Profile:
Professor Inti Zlobec heads the Translational Research Unit at the Institute of Pathology, University of Bern. Her interests lie in inter-disciplinary translational research to improve the clinical management of patients with colorectal cancer and in histomorphological biomarkers as prognostic and predictive features of tumour response to therapies.
26.12.2019





