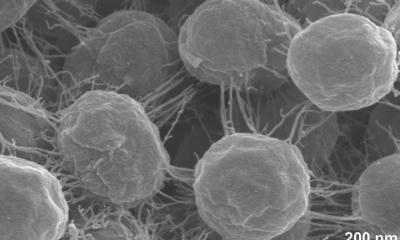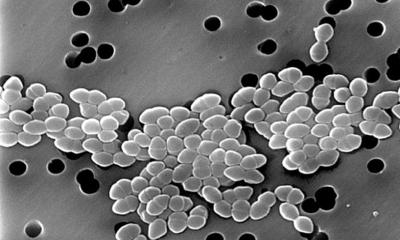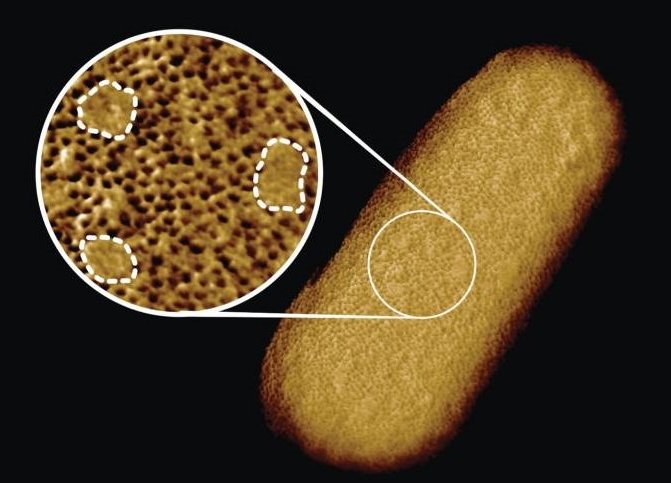
Image credit: Benn et al. UCL
News • Sharper than ever
High-res image of bacterial bumps gives clues to antibiotic resistance
The sharpest images ever of living bacteria have been recorded by researchers at University College London, revealing the complex architecture of the protective layer that surrounds many bacteria and makes them harder to be killed by antibiotics.
The study, published this week in Proceedings of the National Academy of Sciences of the USA and done in collaboration with scientists at National Physical Laboratory, King’s College London, University of Oxford and Princeton University, reveals that bacteria with protective outer layers – called Gram-negative bacteria – may have stronger and weaker spots on their surface.
The team found that the protective outer membrane of the bacteria contains dense networks of protein building blocks alternated by patches that do not appear to contain proteins. Instead, these patches are enriched in molecules with sugary chains (glycolipids) that keep the outer membrane tight.
This new way of looking at the outer membrane means that we can now start exploring if and how such order matters for membrane function, integrity and resistance to antibiotics
Georgina Benn
This is an important finding because the tough outer membrane of Gram-negative bacteria prevents certain drugs and antibiotics from penetrating the cell: this outer membrane is part of the reason why antimicrobial resistance of such bacteria (including A. baumannii, P. aeruginosa, and enterobacteriaceae such as Salmonella and E. coli) is now considered a greater threat than that of Gram-positive bacteria such as resistant S. aureus (well known as MRSA). “The outer membrane is a formidable barrier against antibiotics and is an important factor in making infectious bacteria resistant to medical treatment. However, it remains relatively unclear how this barrier is put together, which is why we chose to study it in such detail,” explained corresponding author Professor Bart Hoogenboom (London Centre for Nanotechnology at UCL and UCL Physics & Astronomy). “By studying live bacteria from the molecular to cellular scale, we can see how membrane proteins form a network that spans the entire surface of the bacteria, leaving small gaps for patches that contain no protein. This suggests that the barrier may not be equally hard to breach or stretch all over the bacterium, but may have stronger and weaker spots that can also be targeted by antibiotics.”
To better understand this architecture, the scientists ran a tiny needle over living Escherichia coli (E. coli) bacteria, thus “feeling” their overall shape. Since the tip of the needle is only a few nanometres wide, this made it possible to visualise molecular structures at the bacterial surface. The resulting images show that the whole outer membrane of the bacteria is crammed with microscopic holes formed by proteins that allow the entry of nutrients while preventing the entry of toxins. Although the outer membrane was known to contain many proteins, this crowded and immobile nature had been unexpected.
Surprisingly, the images also revealed many patches that did not appear to contain proteins. These patches contain a glycolipid normally found on the surface of Gram-negative bacteria. In addition, a different type of pimple-like patch formed when parts of the membrane were flipped inside out due to mutations. In this case, the appearance of these defects correlated with enhanced sensitivity to bacitracin, an antibiotic usually only effective against Gram-positive, but not against Gram-negative bacteria.
As explained by Georgina Benn, who did the microscopy on the bacteria in Professor Hoogenboom’s lab at UCL: “The textbook picture of the bacterial outer membrane shows proteins distributed over the membrane in a disordered manner, well-mixed with other building blocks of the membrane. Our images demonstrate that that is not the case, but that lipid patches are segregated from protein-rich networks just like oil separating from water, in some cases forming chinks in the armour of the bacteria. This new way of looking at the outer membrane means that we can now start exploring if and how such order matters for membrane function, integrity and resistance to antibiotics.”
The team also speculate that the findings may help explain ways by which bacteria can maintain a tightly packed, protective barrier while still allowing rapid growth: the common bacterium E. coli doubles in size and then divides in 20 minutes under favourable conditions. They suggest that the glycolipid patches may allow for more stretch of the membrane than the protein networks, making it easier for the membrane to adapt to the growing size of the bacterium.
Source: University College London
30.10.2021