Acinetobacter baumannii. Scanning electron microscopy.
Image source: © Gudrun Holland; Coloring: Michael Laue/RKI
News • Hospital pathogen Acinetobacter baumannii
More appendages = greater adaptability
Bioinformaticians from Goethe University Frankfurt and Research Unit FOR2251 of the German Research Foundation have now detected an unexpected diversity of certain cell appendages in the bacterium Acinetobacter baumannii that are associated with its pathogenicity.
This could lead to treatment strategies that are specifically tailored to a particular pathogen. The researchers published their findings in the journal PLOS Genetics.
Each year, over 670,000 people in Europe fall ill because of antibiotic-resistant pathogens, and 33,000 die from the infections. Especially feared are pathogens with resistances against multiple, or even all, known antibiotics. One of these is the bacterium Acinetobacter baumannii, feared today above all as the “hospital superbug”: According to estimates, up to five percent of all hospital-acquired and one tenth of all bacterial infections resulting in death can be attributed to this pathogen alone. This puts A. baumannii right at the top of a list of pathogens for which – according to the World Health Organization (WHO) – there is an urgent need to develop new therapies.
Recommended article

Article • Bacterial defense mechanism
Antibiotic resistance: a global threat to healthcare
Antimicrobial resistance (AMR) is becoming more prevalent around the world, constituting a serious threat to public health. When bacteria acquire resistance against antibiotics, common medical procedures – for example, in surgery – become impossible due to the high infection risk. Keep reading to find out about AMR research, development of new antibiotics and antibiotic alternatives.
Understanding which characteristics make A. baumannii a pathogen is one of the prerequisites for this. To this end, bioinformaticians led by Professor Ingo Ebersberger of Goethe University Frankfurt and the LOEWE Center for Translational Biodiversity Genomics (LOEWE-TBG) are comparing the genomes and the proteins encoded therein across a wide range of different Acinetobacter strains. Conclusions about which genes contribute to pathogenicity can be drawn above all from the differences between dangerous and harmless strains.
Due to a lack of suitable methods, corresponding studies have so far concentrated on whether a gene is present in a bacterial strain or not. However, this neglects the fact that bacteria can acquire new characteristics by modifying existing genes and thus also the proteins encoded by them. That is why Ebersberger’s team has developed a bioinformatics method to track the modification of proteins along an evolutionary lineage and has now applied this method for the first time to Acinetobacter in collaboration with microbiologists from the Institute for Molecular Biosciences and the Institute of Medical Microbiology and Infection Control at Goethe University Frankfurt.
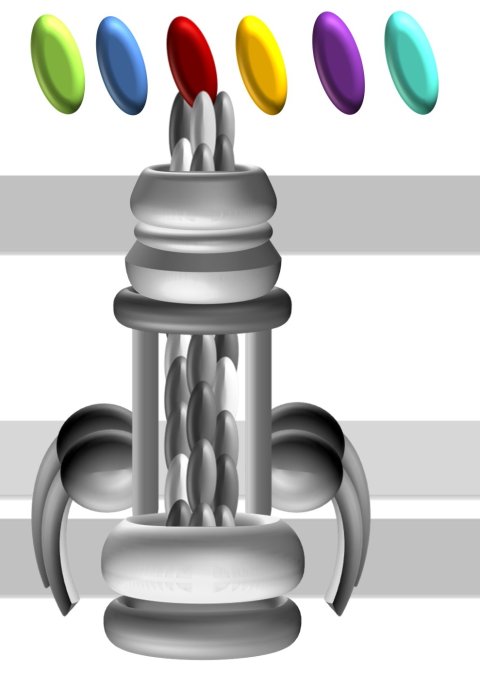

Illustration: Katharina Pfefferle/Goethe University Frankfurt
In the process, the researchers concentrated on hair-like cell appendages, known as type IVa (T4A) pili, which are prevalent in bacteria and that they use to interact with their environment. The fact that they are present in harmless bacteria on the one hand and have even been identified as a key factor for the virulence of some pathogens on the other suggests that the T4A pili have repeatedly acquired new characteristics associated with pathogenicity during evolution.
The research team could show that the protein ComC, which sits on the tip of the T4A pili and is essential for their function, shows conspicuous changes within the group of pathogenic Acinetobacter strains. Even different strains of A. baumannii have different variants of this protein. This leads bioinformatician Ebersberger to compare the T4A pili to a multifunctional garden tool, where the handle is always the same, but the attachments are interchangeable. “In this way, drastic functional modifications can be achieved over short evolutionary time spans,” Ebersberger is convinced. “We assume that bacterial strains that differ in terms of their T4A pili also interact differently with their environment. This might determine, for example, in which corner of the human body the pathogen settles.”
The aim is to use this knowledge of the unexpectedly high diversity within the pathogen to improve the treatment of A. baumannii infections, as Ebersberger explains: “Building on our results, it might be possible to develop personalized therapies that are tailored to a specific strain of the pathogen.” However, the study by Ebersberger and his colleagues also reveals something else: Previous studies on the comparative genomics of A. baumannii have presumably only unveiled the tip of the iceberg. “Our approach has gone a long way towards resolving the search for possible components that characterize pathogens,” says Ebersberger.
Source: Goethe University Frankfurt
04.08.2023